This is a read-only mirror of pymolwiki.org
Difference between revisions of "Gallery"
Jump to navigation
Jump to search
Jaredsampson (talk | contribs) m (fix broken links to Get_View) |
m (1 revision) |
||
| (3 intermediate revisions by 2 users not shown) | |||
| Line 813: | Line 813: | ||
orient org | orient org | ||
ray | ray | ||
| + | </source> | ||
| + | }} | ||
| + | |||
| + | {{GalleryImage | ||
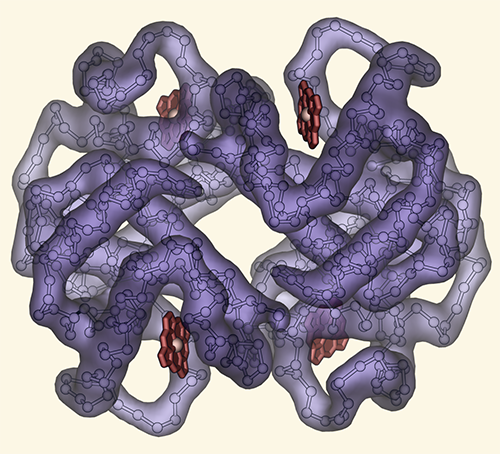
| + | | title=Tribute to Irving Geis | ||
| + | | image=Irvinggeis pymol 500px.png | ||
| + | | description=Hemoglobin, image inspired by Irving Geis | ||
| + | | seeAlso=seeAlso | ||
| + | * [[iterate]] | ||
| + | * [[stick_radius]] | ||
| + | * [[sphere_scale]] | ||
| + | * [[gaussian_b_floor]] | ||
| + | * [[gaussian_resolution]] | ||
| + | * [[map_new]] | ||
| + | * [[isosurface]] | ||
| + | * [[set_color]] | ||
| + | * [[pseudoatom]] | ||
| + | * [[ramp_new]] | ||
| + | * [[cartoon_ring_finder]] | ||
| + | * [[cartoon_ring_mode]] | ||
| + | * [[cartoon_ring_width]] | ||
| + | * [[cartoon_ring_transparency]] | ||
| + | * [[bg_color]] | ||
| + | * [[field_of_view]] | ||
| + | * [[set_view]] | ||
| + | * [[ray_trace_mode]] | ||
| + | * [[ray_shadow]] | ||
| + | * [[light_count]] | ||
| + | * [[light]] | ||
| + | * [[ambient]] | ||
| + | * [[direct]] | ||
| + | * [[specular]] | ||
| + | * [[shininess]] | ||
| + | * [[specular_intensity]] | ||
| + | * [[reflect]] | ||
| + | * [[reflect_power]] | ||
| + | * [[depth_cue]] | ||
| + | * [[fog_start]] | ||
| + | * [[antialias]] | ||
| + | * [[ray]] | ||
| + | | cmdString=<source lang="python"> | ||
| + | # I've always appreciated the simplicity of Irving Geis's designs. | ||
| + | # Here, I've reproduced one of his famous images of hemoglobin. | ||
| + | |||
| + | fetch 1buw | ||
| + | remove solvent | ||
| + | hide all | ||
| + | |||
| + | # Irving Geis shows only the carbonyl carbon atoms in his structure, | ||
| + | # so build a backbone made up of only those atoms | ||
| + | |||
| + | create bbC_A, n. C in /1buw/A/A | ||
| + | stored.bbC = [] | ||
| + | iterate (bbC_A), stored.bbC.append(index) | ||
| + | for i in stored.bbC: cmd.bond("i. %s in bbC_A" % str(i), "i. %s in bbC_A" % str(i+1)) | ||
| + | |||
| + | create bbC_B, n. C in /1buw/B/B | ||
| + | stored.bbC = [] | ||
| + | iterate (bbC_B), stored.bbC.append(index) | ||
| + | for i in stored.bbC: cmd.bond("i. %s in bbC_B" % str(i), "i. %s in bbC_B" % str(i+1)) | ||
| + | |||
| + | create bbC_C, n. C in /1buw/C/C | ||
| + | stored.bbC = [] | ||
| + | iterate (bbC_C), stored.bbC.append(index) | ||
| + | for i in stored.bbC: cmd.bond("i. %s in bbC_C" % str(i), "i. %s in bbC_C" % str(i+1)) | ||
| + | |||
| + | create bbC_D, n. C in /1buw/D/D | ||
| + | stored.bbC = [] | ||
| + | iterate (bbC_D), stored.bbC.append(index) | ||
| + | for i in stored.bbC: cmd.bond("i. %s in bbC_D" % str(i), "i. %s in bbC_D" % str(i+1)) | ||
| + | |||
| + | show sticks, /bbC_A or /bbC_B or /bbC_C or /bbC_D | ||
| + | set stick_radius, 0.2 | ||
| + | show spheres, bbC_A or bbC_B or bbC_C or bbC_D | ||
| + | set sphere_scale, 0.4 | ||
| + | color white, bbC_A or bbC_B or bbC_C or bbC_D | ||
| + | |||
| + | # Create isosurface maps to draw the backbone surface as a tube | ||
| + | |||
| + | set gaussian_b_floor, 40 | ||
| + | set gaussian_resolution, 5 | ||
| + | |||
| + | map_new mapA, gaussian, 1, bbC_A, 60 | ||
| + | isosurface isoA, mapA, 15 | ||
| + | map_new mapB, gaussian, 1, bbC_B, 60 | ||
| + | isosurface isoB, mapB, 15 | ||
| + | map_new mapC, gaussian, 1, bbC_C, 60 | ||
| + | isosurface isoC, mapC, 15 | ||
| + | map_new mapD, gaussian, 1, bbC_D, 60 | ||
| + | isosurface isoD, mapD, 15 | ||
| + | |||
| + | set transparency, 0.2 | ||
| + | |||
| + | # Create a color gradient with a color ramp from a pseudoatom | ||
| + | |||
| + | set_color tubedark, [67, 45, 133] | ||
| + | set_color tubelight, [178, 177, 204] | ||
| + | |||
| + | pseudoatom pseud, pos=[142.982, 26.505, 86.321] | ||
| + | hide /pseud | ||
| + | |||
| + | ramp_new colorRamp, pseud, range=[0,100,142], color=[tubedark, tubedark, tubelight] | ||
| + | set surface_color, colorRamp, isoA | ||
| + | set surface_color, colorRamp, isoB | ||
| + | set surface_color, colorRamp, isoC | ||
| + | set surface_color, colorRamp, isoD | ||
| + | disable colorRamp | ||
| + | |||
| + | # Represent the heme rings as cartoons | ||
| + | |||
| + | create hemes, /1buw/E/A or /1buw/G/B or /1buw/I/C or /1buw/K/D | ||
| + | # Recreate a lost bond... | ||
| + | bond /hemes/K/D/HEM`147/FE, /hemes/K/D/HEM`147/NC | ||
| + | show_as cartoon, hemes | ||
| + | set cartoon_ring_finder, 4 | ||
| + | set cartoon_ring_mode, 3 | ||
| + | set cartoon_ring_width, 0.3 | ||
| + | set cartoon_ring_transparency, 0 | ||
| + | color ruby, hemes | ||
| + | |||
| + | # Represent the heme iron ions as spheres | ||
| + | |||
| + | create irons, e. Fe | ||
| + | show_as spheres, irons | ||
| + | set sphere_scale, 0.6, irons | ||
| + | color darksalmon, irons | ||
| + | |||
| + | # Hide ligands bound to hemes | ||
| + | |||
| + | hide //F or //H or //J or //L | ||
| + | |||
| + | # Hide atoms that stick out of the surface (mostly terminal atoms) | ||
| + | |||
| + | hide /bbC_C/C/C/141 | ||
| + | hide /bbC_A/A/A/141 | ||
| + | hide /bbC_D/D/D/144 | ||
| + | hide /bbC_D/D/D/1 | ||
| + | |||
| + | # Display settings | ||
| + | |||
| + | set_color bground, [252, 247, 229] | ||
| + | bg_color bground | ||
| + | set field_of_view, 5 | ||
| + | set_view (\ | ||
| + | 0.568997085, 0.032208484, 0.821707368,\ | ||
| + | 0.172765702, 0.972248971, -0.157743633,\ | ||
| + | -0.803984761, 0.231720015, 0.547644258,\ | ||
| + | 0.000000000, 0.000000000, -691.255310059,\ | ||
| + | 48.556941986, 45.649765015, 22.998313904,\ | ||
| + | 579.895507812, 802.615112305, 5.000000000 ) | ||
| + | |||
| + | set ray_trace_mode, 1 | ||
| + | set ray_shadow, 0 | ||
| + | set light_count, 2 | ||
| + | set light, [0, 0, -100] | ||
| + | set ambient, 0 | ||
| + | set direct, 0.7 | ||
| + | set specular, 1 | ||
| + | set shininess, 5 | ||
| + | set specular_intensity, 0.3 | ||
| + | set reflect, 0.2 | ||
| + | set reflect_power, 1 | ||
| + | set depth_cue, 1 | ||
| + | set fog_start, 0.45 | ||
| + | set antialias, 3 | ||
| + | |||
| + | ray 1280, 960 | ||
</source> | </source> | ||
}} | }} | ||
Latest revision as of 06:13, 6 March 2017
| Cool PyMOL-generated Images and their Scripts. Add Your Own |
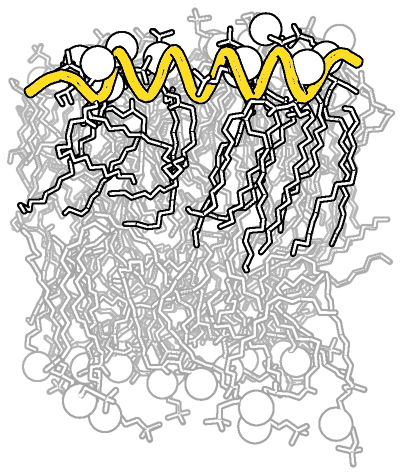
| Complex B&W outline representation | What To Type | |||||
|
# first load lipid model
load lipids.pdb;
# hide the initially loaded representation
hide all;
# set background color to white
bg_color white;
# show lipid model as sticks
show sticks, lipids;
# color the lipids model by element CHNOS #2 (carbon green)
util.cbag lipids;
# select all hydrogens and remove them from the model
select hideme, hydro;
hide everything, hideme;
delete hideme;
# create phosphate spheres
create phos, elem p;
hide everything, phos;
show spheres, phos;
# load helix model
load helix.pdb;
# hide the initially loaded representation
hide everything, helix;
# make the helical struct into a cartoon form
show cartoon, helix;
# style the cartoon form
cartoon putty;
# reposition the helix among the lipids using
# the 3-Button Editing Mouse Mode
# basically
# Shift+Left Mouse to rotate the helix
# Shift+Middle Mouse to move the helix
# also, you may want to make liberal use of the
# get_view and set_view commands.
#
# When you have the scene set like you want,
# continue with...
# move the model to find the view you want,
# and use get_view to get the coordinate description
get_view;
# set ray_trace_mode to black and white outline
set ray_trace_mode, 2;
|
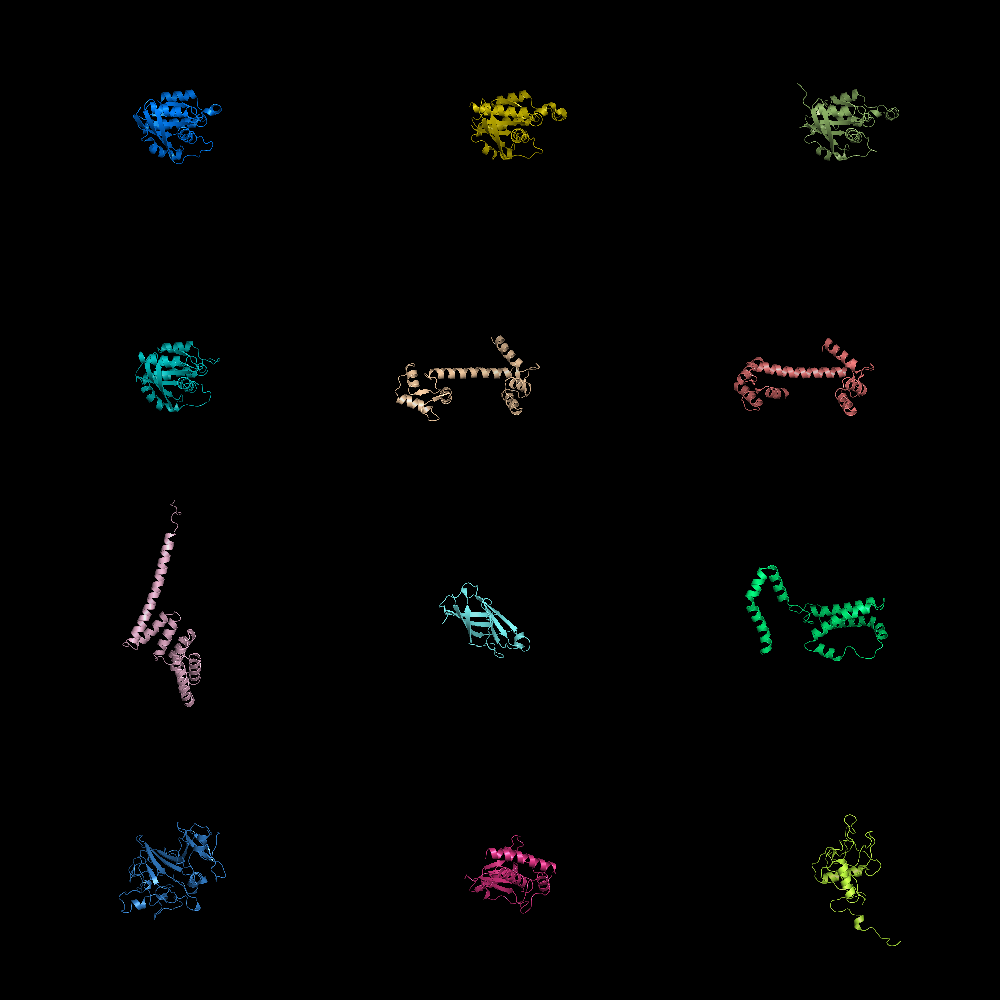
| Grid Mode | What To Type | |||||
|
fetch 1cll 1sra 1ggz 5pnt 1rlw 1cdy;
set grid_mode
Hint: You may wish to execute the 'reset' command on the command line after running the above commands to get full molecules in view of window and centered in a more useable manner. |
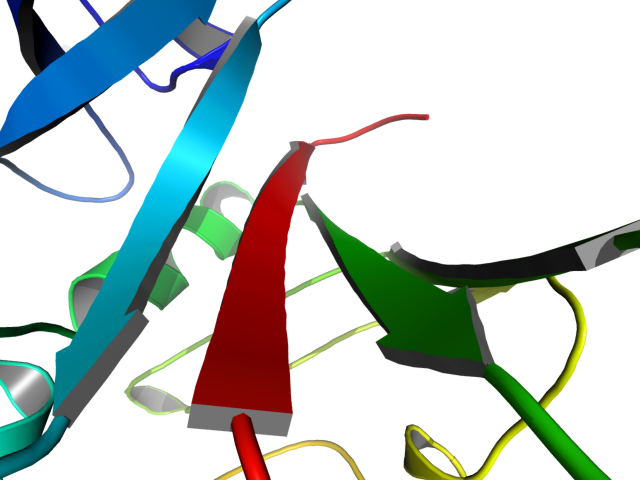
| Cool Perspective | What To Type | |||||
|
load prot.pdb;
zoom i. 46-49 and n. CA
set field_of_view, 60
ray
|
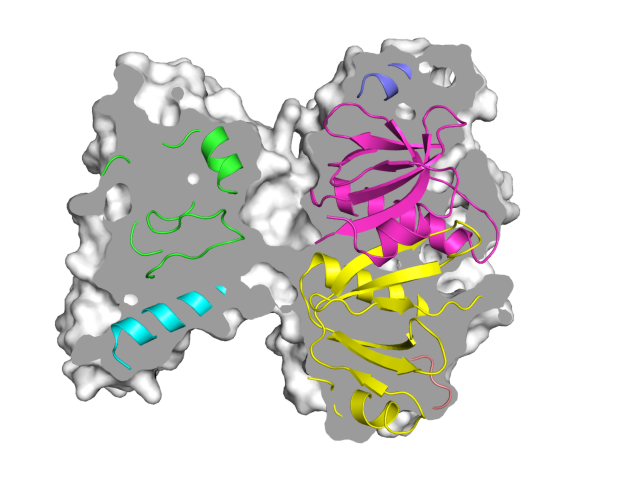
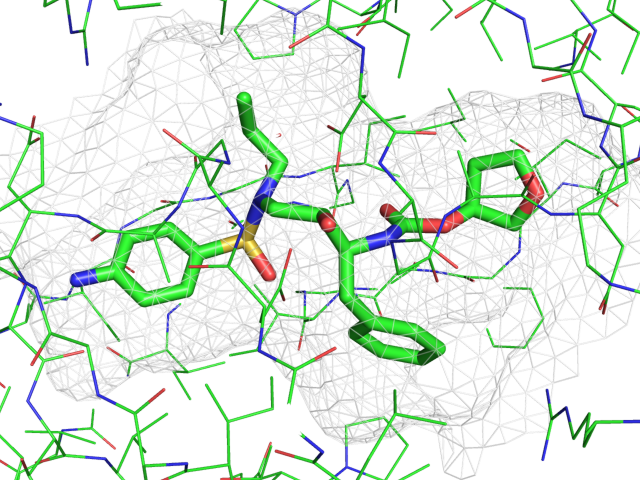
| Representing a binding pocket | What To Type | |||||
|
load $TUT/1hpv.pdb, tmp
extract lig, organic
extract prot, polymer
delete tmp
set surface_carve_cutoff, 4.5
set surface_carve_selection, lig
set surface_carve_normal_cutoff, -0.1
show surface, prot within 8 of lig
set two_sided_lighting
set transparency, 0.5
show sticks, lig
orient lig
set surface_color, white
set surface_type, 2 # mesh
unset ray_shadows
|
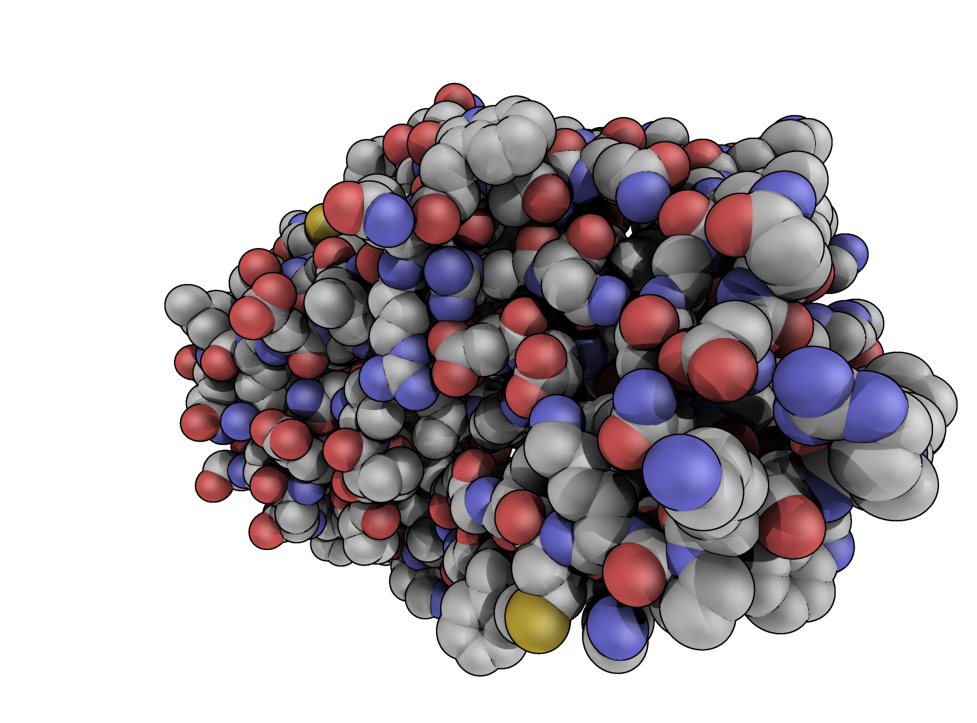
| QuteMol Like | What To Type | |||||
|
load $TUT/1hpv.pdb
set_color oxygen, [1.0,0.4,0.4]
set_color nitrogen, [0.5,0.5,1.0]
remove solvent
as spheres
util.cbaw
bg white
set light_count,8
set spec_count,1
set shininess, 10
set specular, 0.25
set ambient,0
set direct,0
set reflect,1.5
set ray_shadow_decay_factor, 0.1
set ray_shadow_decay_range, 2
unset depth_cue
# for added coolness
# set field_of_view, 60
ray
|
| Simulating Tilt-shift | What To Type | |||||
|
fetch 1wld
as surface, poly
as sticks, org
h_add solvent
color grey, poly
orient org
png img.png
# now, go into Photoshop or the GIMP and apply a Gaussian or
# Focus blur to the top and bottom portions of the image
|
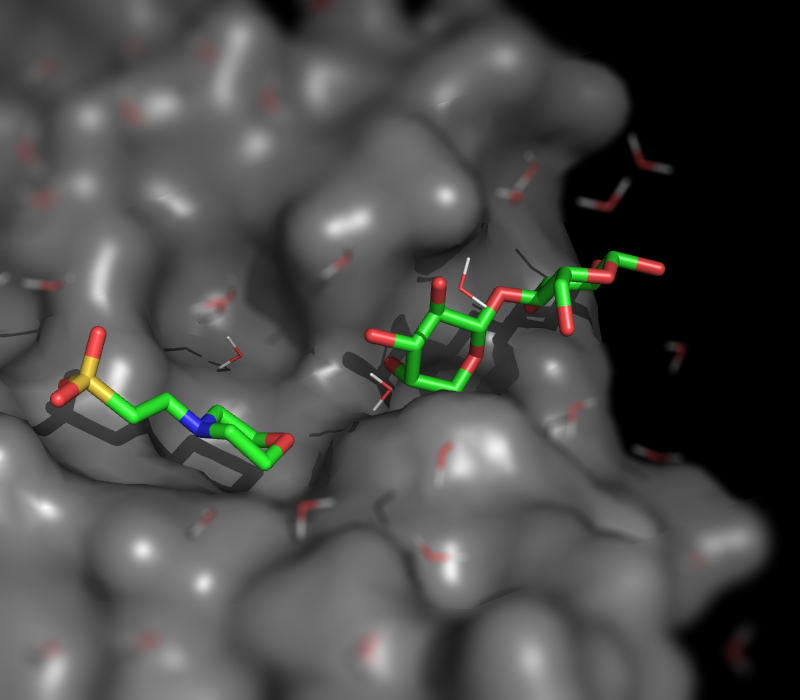
| Ray-normal-based transparency | What To Type | |||||
|
# grey surface
set surface_color, grey
# cavity mode
set surface_mode, 3
# layered transparency mode
set transparency_mode, 1
# surface transparency
set transparency, 0.5
# oblique and contrast define the
# look of the surface transparency:
# if the normal vector is
set ray_transparency_oblique
set ray_transparency_oblique_power, 8
set ray_transparency_contrast, 7
# fetch a protein, with a
# small molecule in a nice
# hidden pocket
fetch 1hpv, async=0
hide
# show the small molecule as surface
show surface, org
# arrange the view
orient org
# zoom back a little
zoom org, 1
# show the small molecule inside as sticks
show sticks, org
# show some nearby sidechains
show lines, poly within 5 of org
# enable frame caching for playback
set cache_frames, 1
set ray_trace_frames, 1
mset 1x120
movie.roll 1, 120, 1, x
mplay
# now sit back and wait 5 minutes...
|
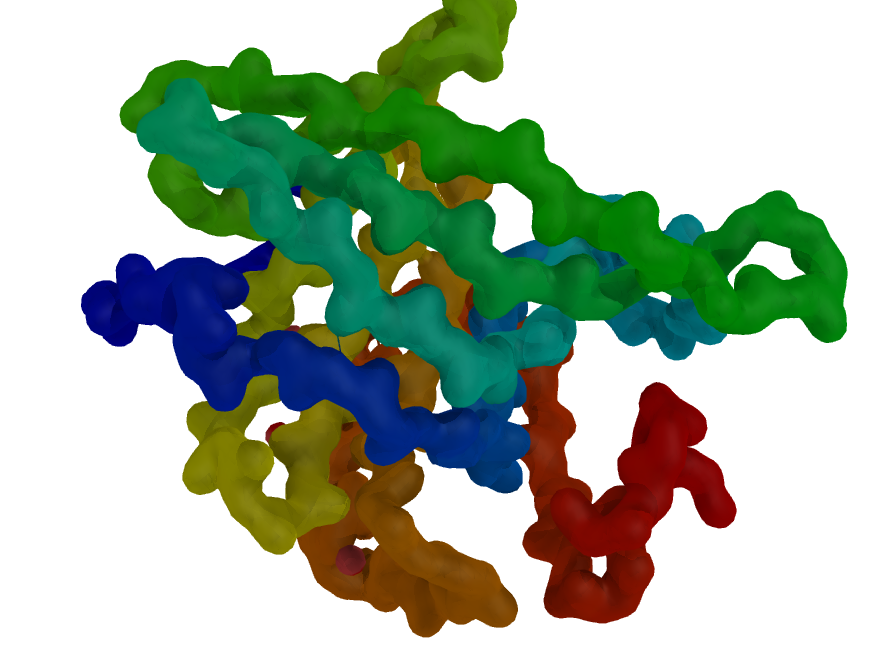
| Blobby Main Chain | What To Type | |||||
|
fetch 3uex, struct, async=0
remove solvent
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 3.5 A map resolution
set gaussian_resolution, 3.5
# new gaussian map w/resolution=0.5 Ang
# on just the main chain
map_new map, gaussian, 0.5, n. C+O+N+CA, 5
# create a surface from the map
isosurface surf, map, 3.0
# color the protein by number
spectrum count, rainbow, struct
# now color the map based on the underlying protein
cmd.ramp_new("ramp", "struct", [0,10,10], [-1, -1, 0])
# set the surface color
cmd.set("surface_color", "ramp", "surf")
# hide the ramp and lines
disable ramp
hide lines
bg grey
# soften out the image
set light_count,8
set spec_count,1
set shininess, 10
set specular, 0.075
set ambient,0
set direct,0
set reflect, 0.85
set ray_shadow_decay_factor, 0.1
set ray_shadow_decay_range, 4
unset depth_cue
# ray trace the image
orient
ray
|
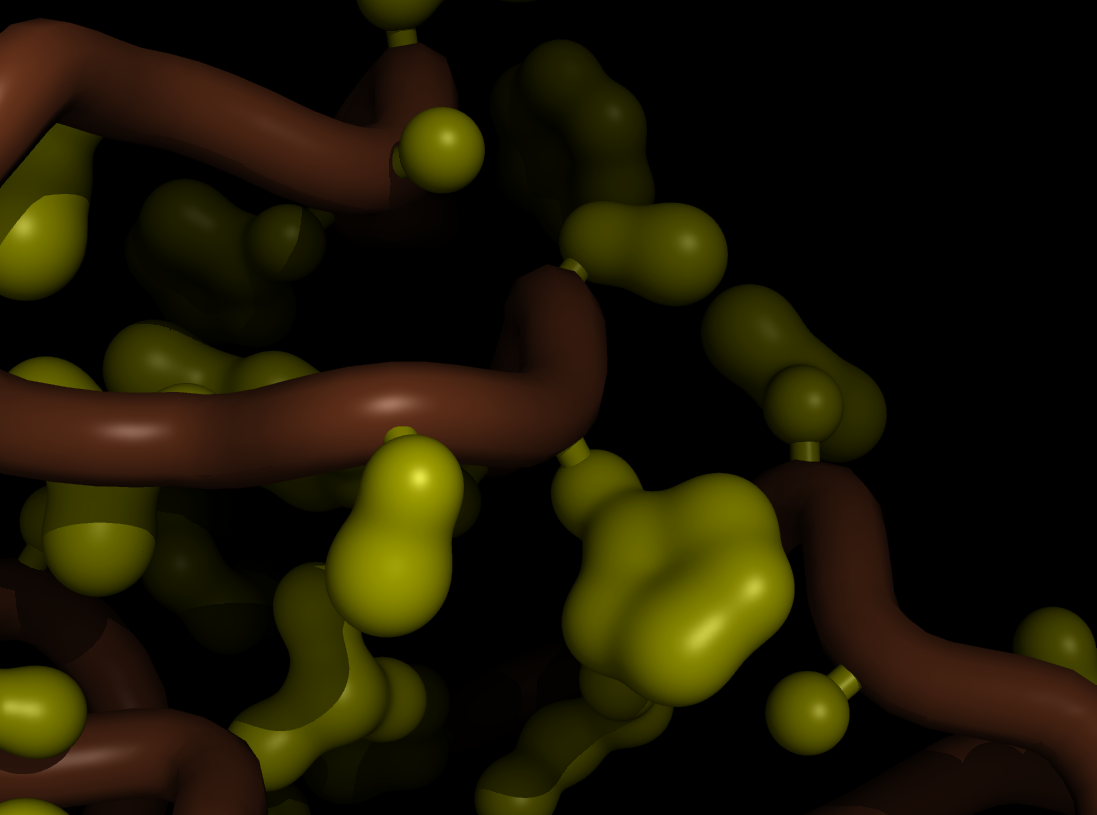
| Blobby Side Chains | What To Type | |||||
|
# fetch a protein
fetch 1rx1, async=0
# setup the cartoon tubes
as cartoon
cartoon tube
set cartoon_tube_radius, 0.7
set cartoon_color, brown
set cartoon_side_chain_helper, on
show sticks, poly
color yellow
# README
# stop here, or try this for "sloppy sticks"
# beefy video card required!
select rep sticks
select sele and not n. C+O+N+CA
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 2.5 A map resolution
set gaussian_resolution, 2.5
# 0.2 A sampling; lower=smoother
map_new map, gaussian, 0.2, sele, 5
# create a surface from the map
isosurface surf, map, 5.0
color yellow, surf
hide sticks
# reconnect the main chain to the blobs
show sticks, n. CA+CB
|
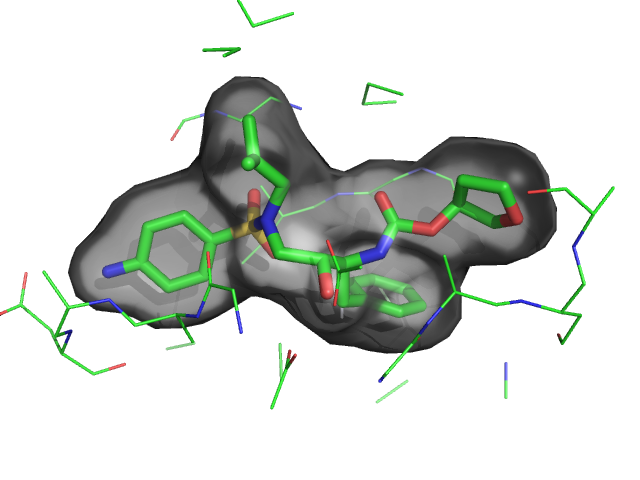
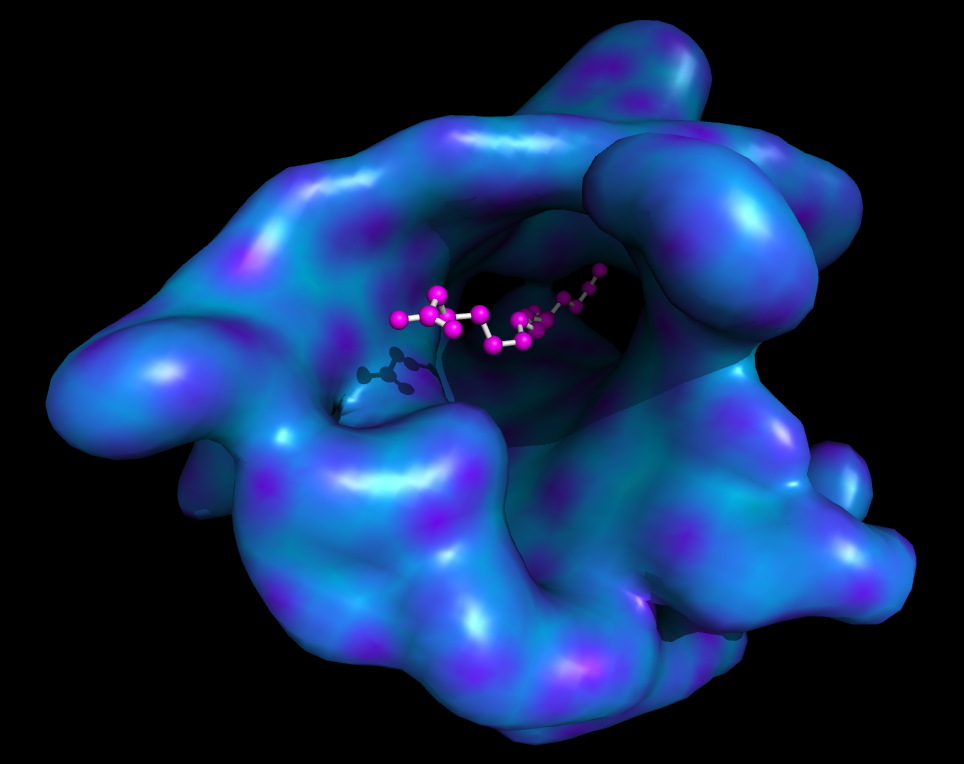
| Smooth surface with ligand | What To Type | |||||
|
fetch 3uex, struct, async=0
remove solvent
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 3.5 A map resolution
set gaussian_resolution, 7.6
# new gaussian map w/resolution=0.5 Ang
# on just the main chain
map_new map, gaussian, 1, n. C+O+N+CA, 5
# create a surface from the map
isosurface surf, map, 1.5
# color the protein by number
spectrum count, rainbow, struct
# now color the map based on the b-factors of the
# underlying protein
cmd.ramp_new("ramp", "struct", [0,10,10], "rainbow")
# set the surface color
cmd.set("surface_color", "ramp", "surf")
# hide the ramp and lines
disable ramp
hide lines
show sticks, org
show spheres, org
color magenta, org
reinit
fetch 3uex, struct, async=0
remove solvent
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 3.5 A map resolution
set gaussian_resolution, 7.6
# new gaussian map w/resolution=0.5 Ang
# on just the main chain
map_new map, gaussian, 1, n. C+O+N+CA, 5
# create a surface from the map
isosurface surf, map, 1.5
# color the protein by number
spectrum count, rainbow, struct
# now color the map based on the b-factors of the
# underlying protein
cmd.ramp_new("ramp", "struct", [0,10,10], "rainbow")
# set the surface color
cmd.set("surface_color", "ramp", "surf")
# hide the ramp and lines
disable ramp
hide lines
show sticks, org
show spheres, org
color magenta, org
set_bond stick_radius, 0.13, org
set sphere_scale, 0.26, org
set_bond stick_radius, 0.13, org
set_bond stick_color, white, org
set sphere_scale, 0.26, org
set_view (\
-0.877680123, 0.456324875, -0.146428943,\
0.149618521, -0.029365506, -0.988305628,\
-0.455291569, -0.889327347, -0.042500813,\
-0.000035629, 0.000030629, -37.112102509,\
-3.300258160, 6.586110592, 22.637466431,\
8.231912613, 65.999290466, -50.000000000 )
# ray trace the image
ray
|
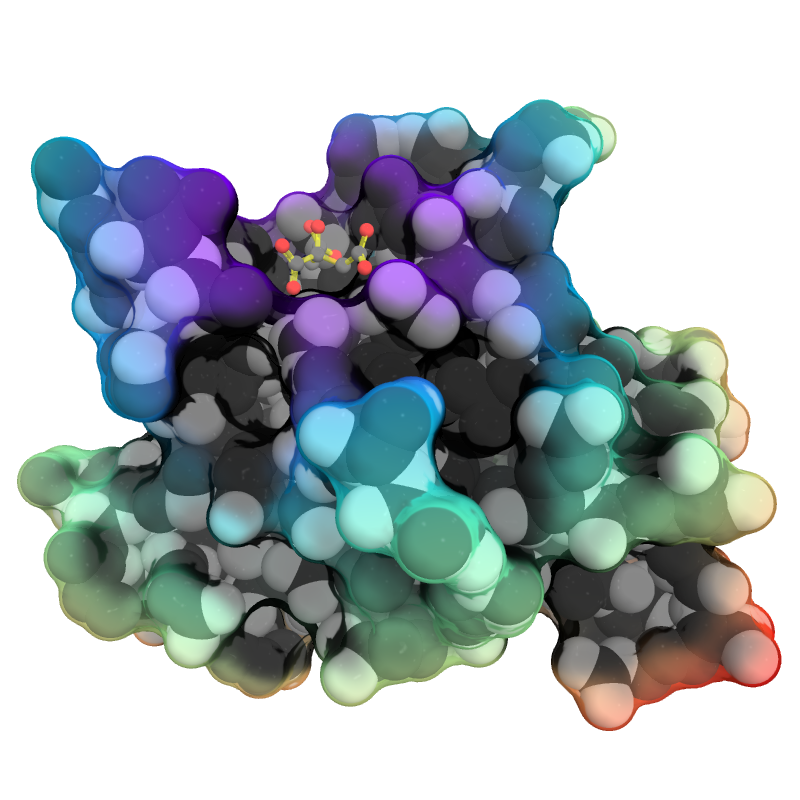
| Complex Stylized Protein | What To Type | |||||
|
fetch 1eaz, async=0
extract oo, org
hide everything, solvent
set field_of_view, 50
preset.ball_and_stick("oo")
set_bond stick_color, 0xffff44, oo
set_bond stick_transparency, 0.35, oo
color grey, oo and e. C
set valence, 1, oo
ramp_new pRamp, oo, selection=poly, range=[5,30], color=rainbow
set surface_color, pRamp, poly
show spheres, poly
color white, poly
color grey30, poly and e. C
set sphere_scale, 0.99, poly
set ray_transparency_contrast, 0.20
set ray_transparency_oblique, 1.0
set ray_transparency_oblique_power, 20
show surface, poly
set surface_quality, 2
set light_count, 5
set ambient_occlusion_mode, 1
set ambient_occlusion_scale, 50
set ambient, 0.40
set transparency, 0.50
disable pRamp
set spec_power, 1200
set spec_reflect, 0.20
set ray_opaque_background, 0
set ray_shadow, 0
ray
|
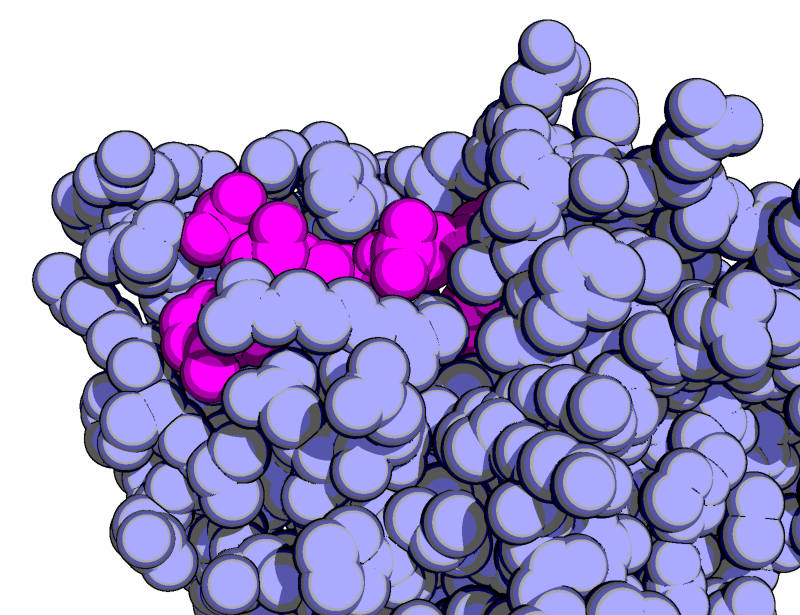
| Goodsell-like | What To Type | |||||
|
# fetch the protein
fetch 1rx1, async=0
# show it as blue/magenta spheres
as spheres
color lightblue, not org
color magenta, org
remove solvent
# set the view
orient all within 8 of org
# set the lights, ray tracing setttings
# to get the Goodsell-like rendering
unset specular
set ray_trace_gain, 0
set ray_trace_mode, 3
bg_color white
set ray_trace_color, black
unset depth_cue
ray
|
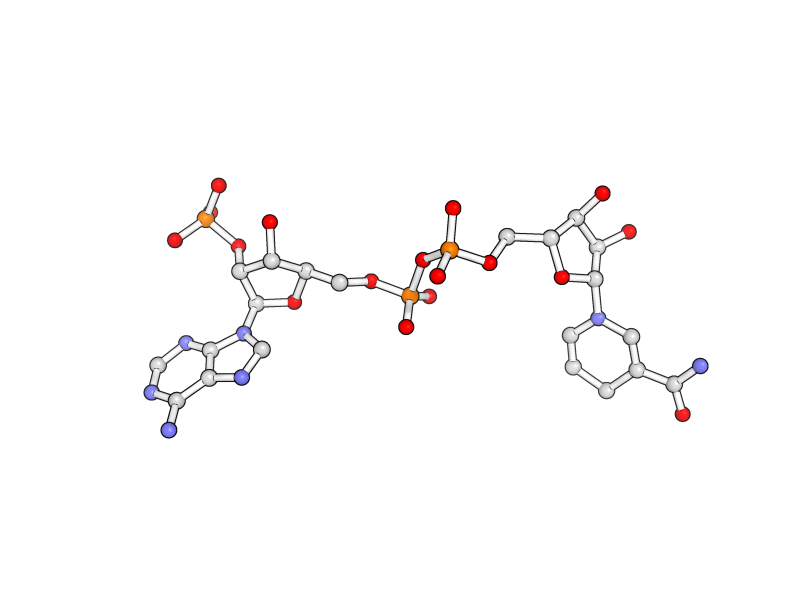
| Stylized Ball and Stick | What To Type | |||||
|
hide everything
show sticks
show spheres
set stick_radius, .07
set sphere_scale, .18
set sphere_scale, .13, elem H
set bg_rgb=[1, 1, 1]
set stick_quality, 50
set sphere_quality, 4
color gray85, elem C
color red, elem O
color slate, elem N
color gray98, elem H
set stick_color, black
set ray_trace_mode, 1
set ray_texture, 2
set antialias, 3
set ambient, 0.5
set spec_count, 5
set shininess, 50
set specular, 1
set reflect, .1
set dash_gap, 0
set dash_color, black
set dash_gap, .15
set dash_length, .05
set dash_round_ends, 0
set dash_radius, .05
python
preset.ball_and_stick("vis")
python end
fetch 1rx1, async=0
remove not org
orient org
ray
|
| Tribute to Irving Geis | What To Type | |||||
|
# I've always appreciated the simplicity of Irving Geis's designs.
# Here, I've reproduced one of his famous images of hemoglobin.
fetch 1buw
remove solvent
hide all
# Irving Geis shows only the carbonyl carbon atoms in his structure,
# so build a backbone made up of only those atoms
create bbC_A, n. C in /1buw/A/A
stored.bbC = []
iterate (bbC_A), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_A" % str(i), "i. %s in bbC_A" % str(i+1))
create bbC_B, n. C in /1buw/B/B
stored.bbC = []
iterate (bbC_B), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_B" % str(i), "i. %s in bbC_B" % str(i+1))
create bbC_C, n. C in /1buw/C/C
stored.bbC = []
iterate (bbC_C), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_C" % str(i), "i. %s in bbC_C" % str(i+1))
create bbC_D, n. C in /1buw/D/D
stored.bbC = []
iterate (bbC_D), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_D" % str(i), "i. %s in bbC_D" % str(i+1))
show sticks, /bbC_A or /bbC_B or /bbC_C or /bbC_D
set stick_radius, 0.2
show spheres, bbC_A or bbC_B or bbC_C or bbC_D
set sphere_scale, 0.4
color white, bbC_A or bbC_B or bbC_C or bbC_D
# Create isosurface maps to draw the backbone surface as a tube
set gaussian_b_floor, 40
set gaussian_resolution, 5
map_new mapA, gaussian, 1, bbC_A, 60
isosurface isoA, mapA, 15
map_new mapB, gaussian, 1, bbC_B, 60
isosurface isoB, mapB, 15
map_new mapC, gaussian, 1, bbC_C, 60
isosurface isoC, mapC, 15
map_new mapD, gaussian, 1, bbC_D, 60
isosurface isoD, mapD, 15
set transparency, 0.2
# Create a color gradient with a color ramp from a pseudoatom
set_color tubedark, [67, 45, 133]
set_color tubelight, [178, 177, 204]
pseudoatom pseud, pos=[142.982, 26.505, 86.321]
hide /pseud
ramp_new colorRamp, pseud, range=[0,100,142], color=[tubedark, tubedark, tubelight]
set surface_color, colorRamp, isoA
set surface_color, colorRamp, isoB
set surface_color, colorRamp, isoC
set surface_color, colorRamp, isoD
disable colorRamp
# Represent the heme rings as cartoons
create hemes, /1buw/E/A or /1buw/G/B or /1buw/I/C or /1buw/K/D
# Recreate a lost bond...
bond /hemes/K/D/HEM`147/FE, /hemes/K/D/HEM`147/NC
show_as cartoon, hemes
set cartoon_ring_finder, 4
set cartoon_ring_mode, 3
set cartoon_ring_width, 0.3
set cartoon_ring_transparency, 0
color ruby, hemes
# Represent the heme iron ions as spheres
create irons, e. Fe
show_as spheres, irons
set sphere_scale, 0.6, irons
color darksalmon, irons
# Hide ligands bound to hemes
hide //F or //H or //J or //L
# Hide atoms that stick out of the surface (mostly terminal atoms)
hide /bbC_C/C/C/141
hide /bbC_A/A/A/141
hide /bbC_D/D/D/144
hide /bbC_D/D/D/1
# Display settings
set_color bground, [252, 247, 229]
bg_color bground
set field_of_view, 5
set_view (\
0.568997085, 0.032208484, 0.821707368,\
0.172765702, 0.972248971, -0.157743633,\
-0.803984761, 0.231720015, 0.547644258,\
0.000000000, 0.000000000, -691.255310059,\
48.556941986, 45.649765015, 22.998313904,\
579.895507812, 802.615112305, 5.000000000 )
set ray_trace_mode, 1
set ray_shadow, 0
set light_count, 2
set light, [0, 0, -100]
set ambient, 0
set direct, 0.7
set specular, 1
set shininess, 5
set specular_intensity, 0.3
set reflect, 0.2
set reflect_power, 1
set depth_cue, 1
set fog_start, 0.45
set antialias, 3
ray 1280, 960
|