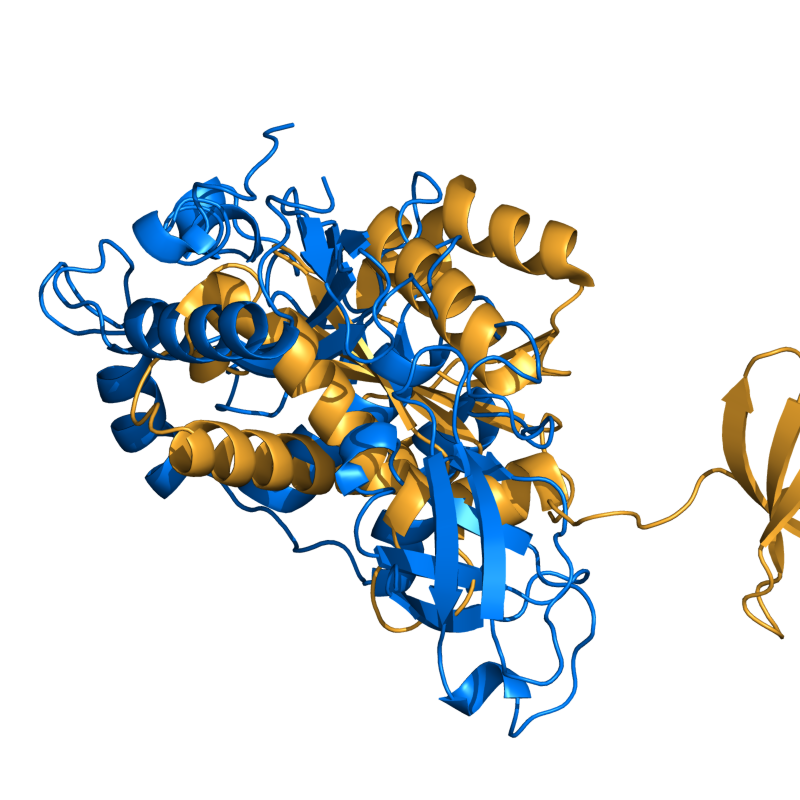This is a read-only mirror of pymolwiki.org
Uncategorized files
Jump to navigation
Jump to search
Showing below up to 500 results in range #251 to #750.
View (previous 500 | next 500) (20 | 50 | 100 | 250 | 500)
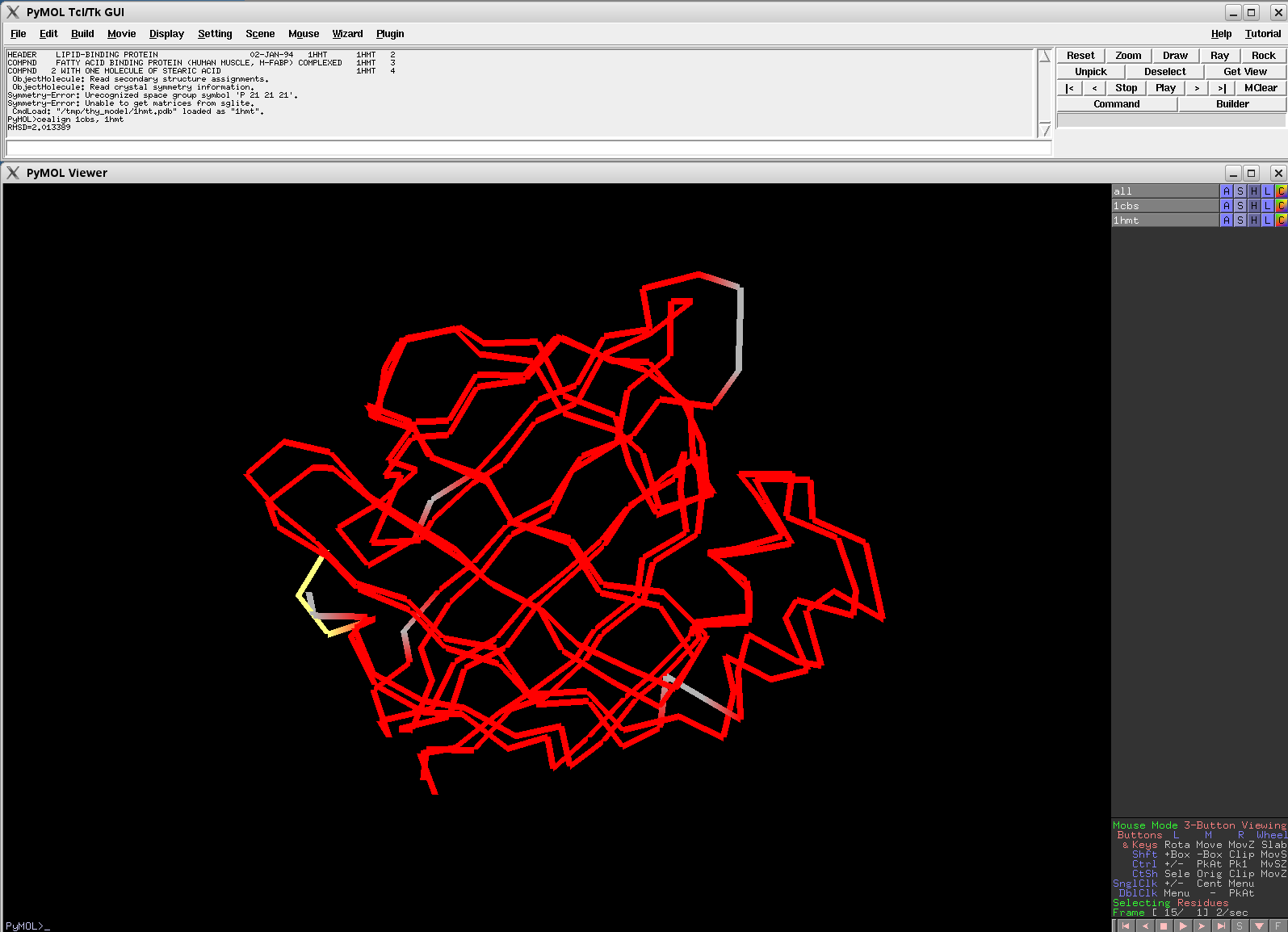
- Cealign2.png 1,595 × 1,154; 52 KB
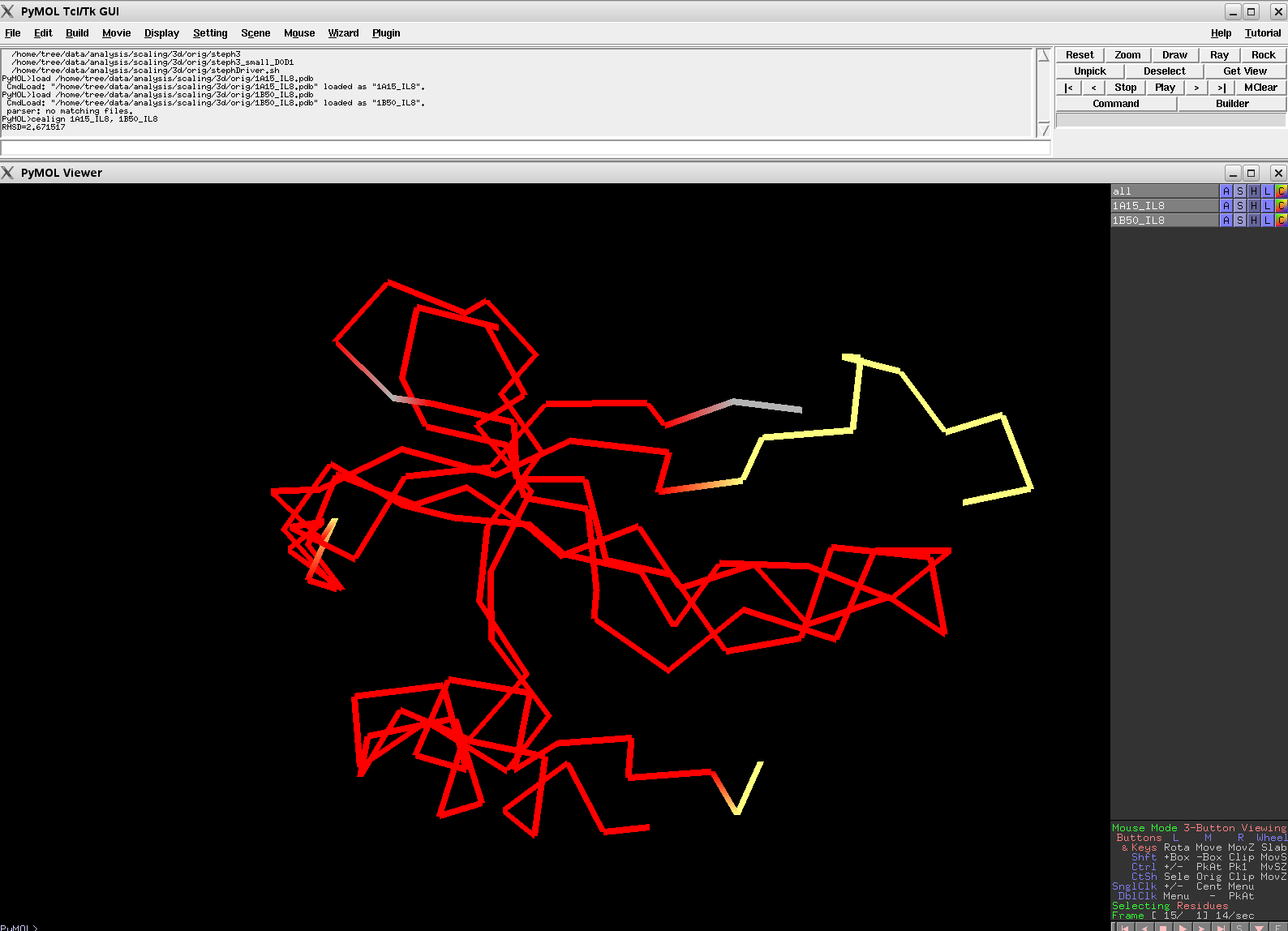
- Cealign3.png 1,588 × 1,148; 46 KB
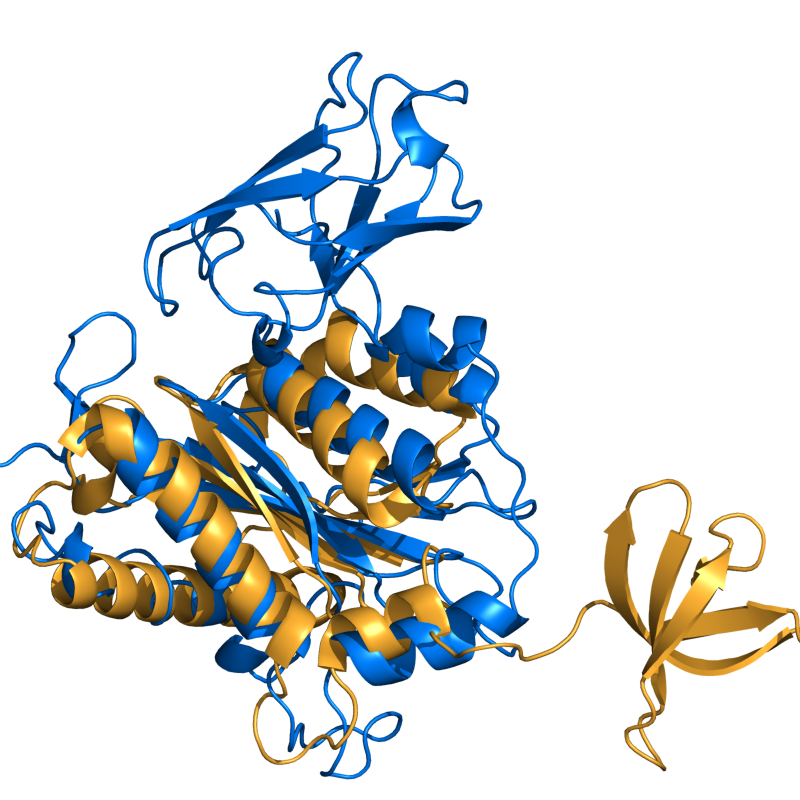
- Cealign ex1.png 800 × 800; 359 KB
- Cealign ex hard.png 600 × 400; 132 KB
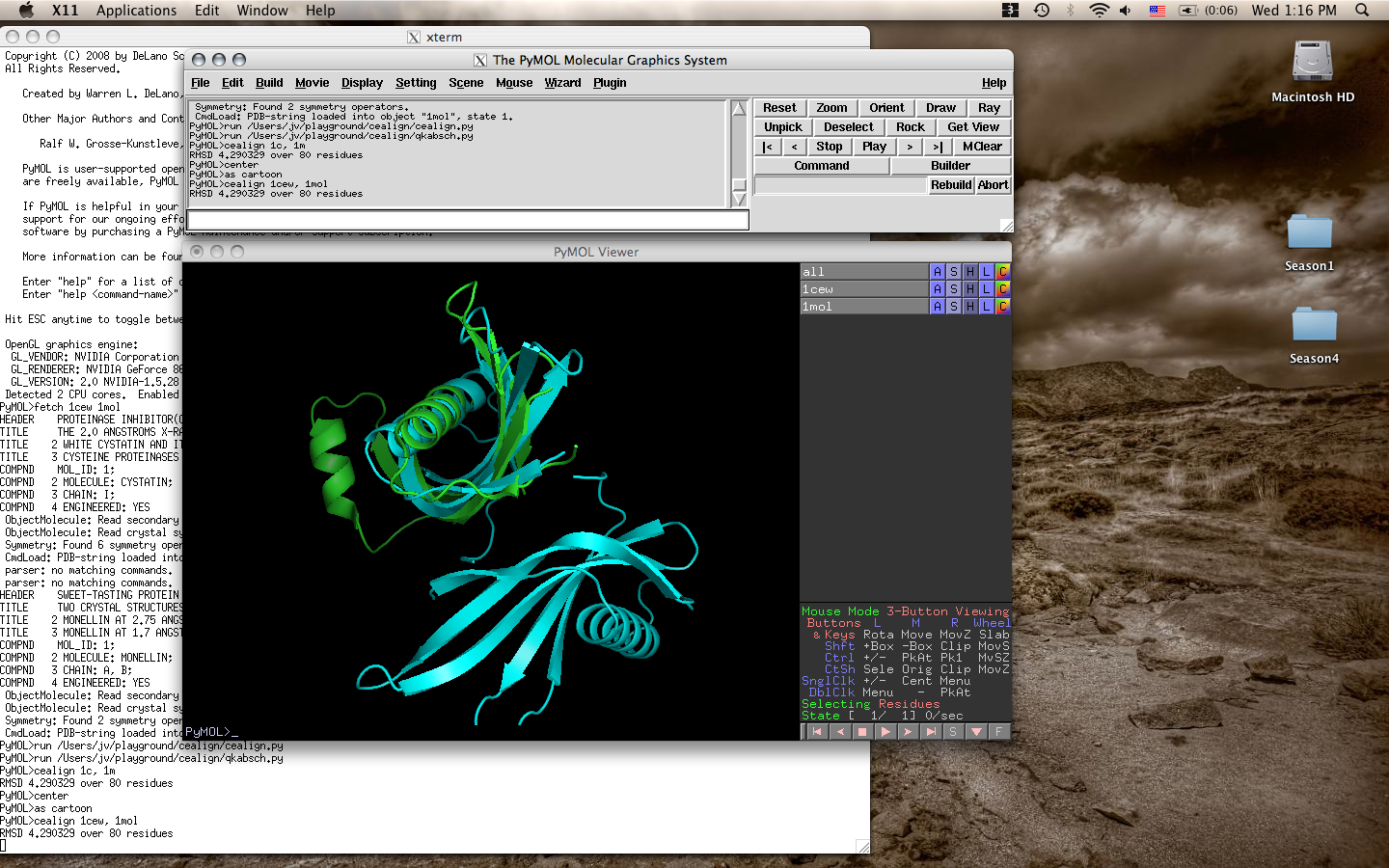
- Cealign mac os x.png 1,440 × 900; 668 KB
- Cen.jpg 640 × 808; 112 KB
- Cfs off.png 400 × 400; 82 KB
- Cfs on.png 400 × 400; 66 KB
- Cgo arrow example.png 400 × 300; 34 KB
- Cgo grid.gif 300 × 200; 870 KB
- Cgo grid.png 930 × 412; 28 KB
- Cgo text.png 843 × 720; 54 KB
- Cheshift.png 1,305 × 723; 133 KB
- Chroma 3d.png 1,288 × 874; 623 KB
- Chroma 3d2.png 1,288 × 874; 304 KB
- Chroma 3d3.png 1,250 × 944; 665 KB
- Chroma 3d4.png 1,250 × 944; 663 KB
- Chroma 3d5.png 1,250 × 944; 170 KB
- Circle1.png 1,399 × 919; 76 KB
- Circle2.png 1,399 × 919; 589 KB
- Circle3.png 1,399 × 919; 268 KB
- CircleR.png 640 × 480; 211 KB
- CircleR2.png 640 × 480; 323 KB
- Close ray.png 300 × 300; 59 KB
- Cls.png 870 × 526; 21 KB
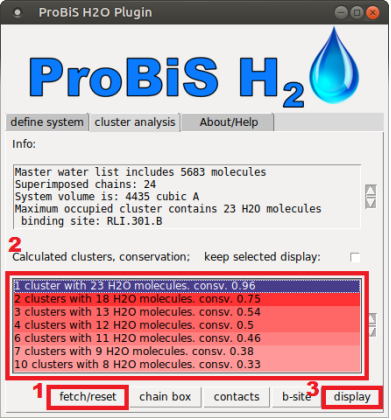
- Cluster analysis tab.PNG 391 × 420; 102 KB
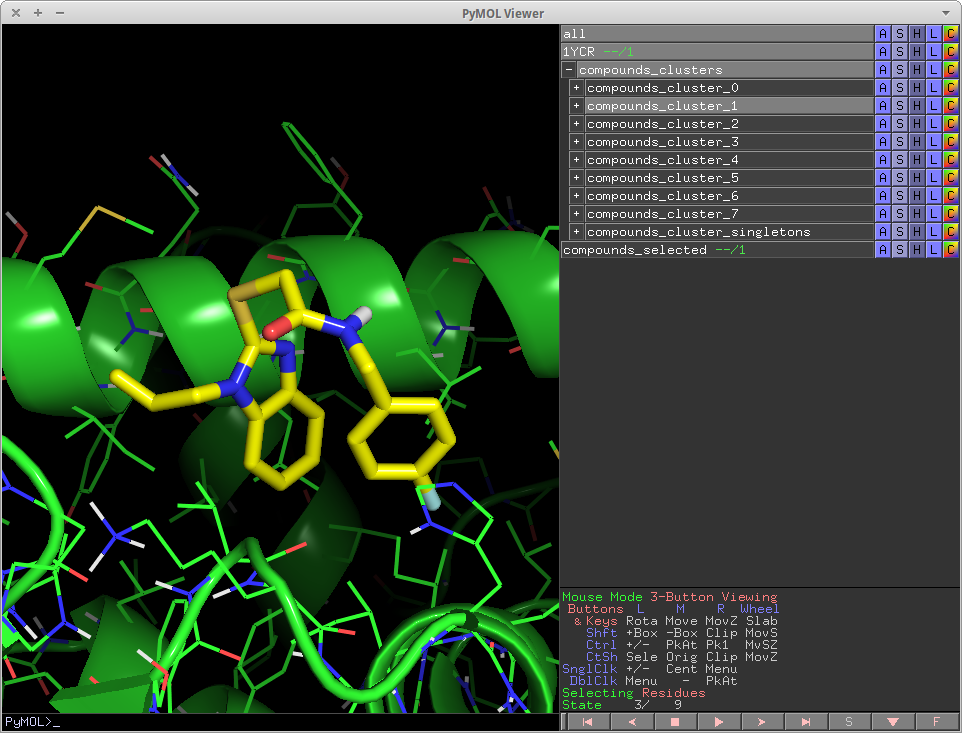
- Cluster mols py pymol.png 962 × 733; 220 KB
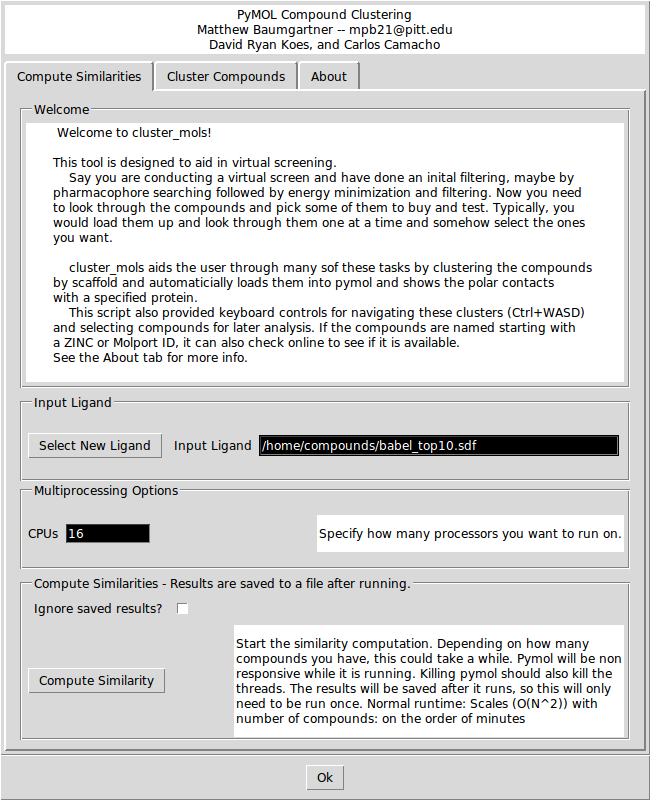
- Cluster mols screen 1 desc.png 650 × 800; 66 KB
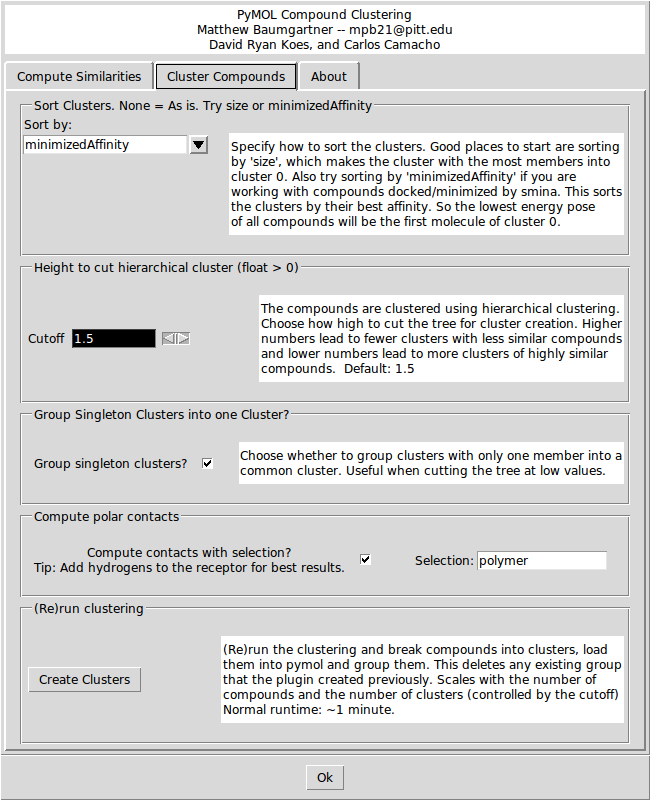
- Cluster mols screen 2 desc.png 650 × 800; 67 KB
- Cna color1.png 640 × 480; 130 KB
- Cna color2.png 640 × 480; 96 KB
- Cna color3.pngCna color3.png File missing
- Cnam 0.png 640 × 480; 153 KB
- Cnam 1.png 640 × 480; 145 KB
- Cnam 2.png 640 × 480; 155 KB
- Cns.png 400 × 300; 72 KB
- Code search links.png 984 × 485; 89 KB
- ColorByDisplacement-All-1.png 640 × 480; 174 KB
- ColorByDisplacement-All-2.png 640 × 480; 152 KB
- ColorByDisplacement-CA-1.png 640 × 480; 110 KB
- ColorByDisplacement-CA-2.png 640 × 480; 98 KB
- ColorByRMSD 1cbs 1hmt.png 707 × 513; 127 KB
- ColorByRMSD 1eaz 1fao.png 707 × 513; 112 KB
- Color by mutation.png 640 × 480; 162 KB
- Colorama-0.1.0.tar.bz2Colorama-0.1.0.tar.bz2 File missing
- Colorama-0.1.1.tar.bz2Colorama-0.1.1.tar.bz2 File missing
- Colorama.tar.bz2Colorama.tar.bz2 File missing
- Colorbar-blue green.png 1,058 × 100; 4 KB
- Colorblindfriendly menu.png 578 × 954; 42 KB
- Connect cutoff.png 339 × 480; 32 KB
- ConnectedCloud.png 800 × 800; 547 KB
- ConnectedCloud2.png 800 × 800; 545 KB
- Contact-Map-of-a-Trajectory.png 308 × 308; 29 KB
- Contact density.png 1,252 × 675; 38 KB
- Cool.png 800 × 800; 481 KB
- Cover JCIM 2008 48-10.png 639 × 850; 257 KB
- Cover Paunescu.jpg 2,480 × 3,307; 2.22 MB
- Cover royal soc small.png 600 × 915; 640 KB
- Crm 0.png 640 × 480; 109 KB
- Crm 1.png 640 × 480; 137 KB
- Crm 2.png 640 × 480; 130 KB
- Crm 3.png 640 × 480; 185 KB
- Crm 4.png 640 × 480; 171 KB
- Crm 6.png 640 × 480; 130 KB
- Crt 05.png 640 × 480; 190 KB
- Crt 1.png 640 × 480; 207 KB
- Csl off.png 400 × 400; 58 KB
- Csl on.png 400 × 400; 40 KB
- Csos3-stereo01.png 2,160 × 1,500; 1.18 MB
- Ctt.png 640 × 480; 187 KB
- Cull backface off.jpg 320 × 240; 50 KB
- Cull backface on.jpg 320 × 240; 49 KB
- CylCGO.png 600 × 400; 46 KB
- Cylindre.JPG 182 × 204; 4 KB
- Cylsphere.png 600 × 400; 33 KB
- Cyspka.png 500 × 500; 113 KB
- DL0 10.png 640 × 480; 4 KB
- DL0 15.png 640 × 480; 4 KB
- DL0 50.png 640 × 480; 3 KB
- DL4.png 640 × 480; 3 KB
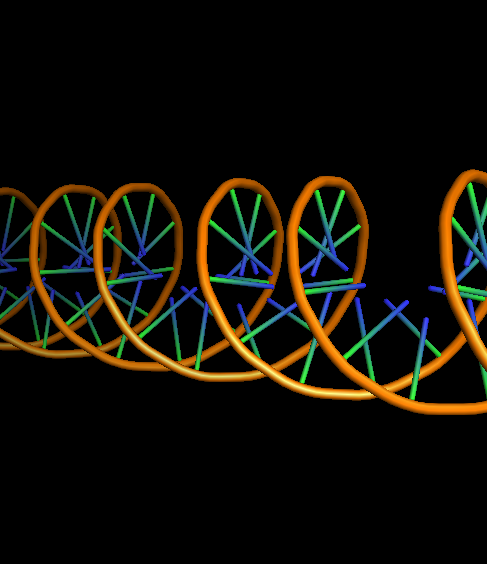
- DNA-default-ring0-ladder1-na0-finder1.png 400 × 400; 93 KB
- DNA-ring0-ladder1-na0-finder1.png 400 × 400; 93 KB
- DNA-ring1-ladder1-na0-finder1.png 400 × 400; 93 KB
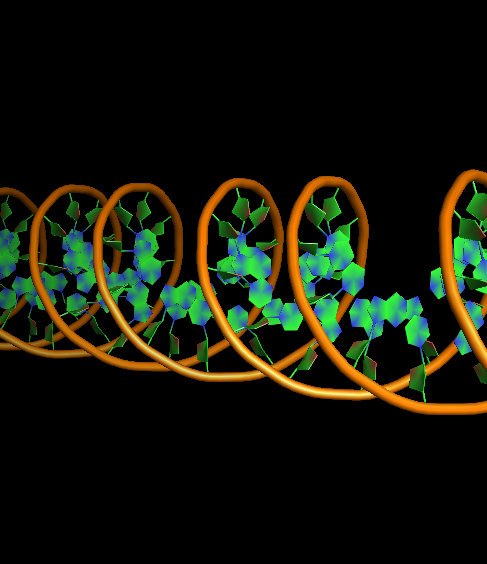
- DNA-ring2-ladder1-na0-finder1.png 400 × 400; 90 KB
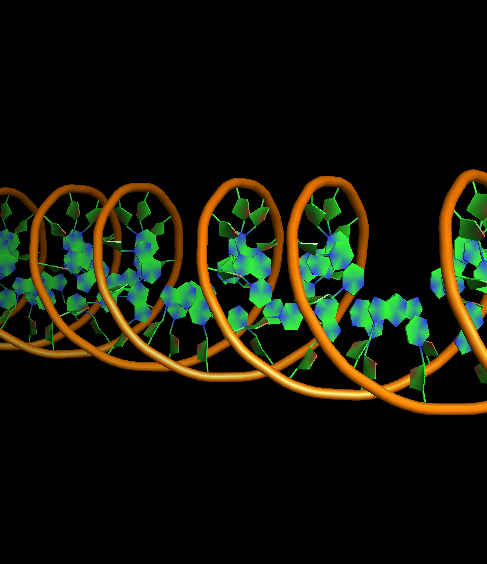
- DNA-ring3-ladder0-na0-finder1.png 400 × 400; 105 KB
- DNA-ring3-ladder1-na0-finder0.png 400 × 400; 48 KB
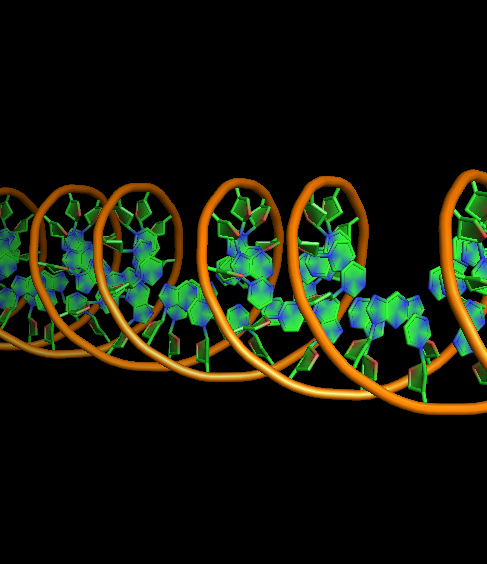
- DNA-ring3-ladder1-na0-finder1.png 400 × 400; 110 KB
- DNA-ring3-ladder1-na0-finder2.png 400 × 400; 101 KB
- DNA-ring3-ladder1-na0-finder3.png 400 × 400; 110 KB
- DNA-ring3-ladder1-na0-finder4.png 400 × 400; 110 KB
- DNA-ring3-ladder1-na0-finder5.png 400 × 400; 66 KB
- DNA-ring3-ladder1-na1-finder1.png 400 × 400; 98 KB
- DNA-ring3-ladder1-na2-finder1.png 400 × 400; 110 KB
- DNA-ring3-ladder1-na3-finder1.png 400 × 400; 110 KB
- DNA-ring3-ladder1-na4-finder1.png 400 × 400; 110 KB
- DNA-ring4-ladder1-na0-finder1.png 400 × 400; 106 KB
- DNA-ring5-ladder1-na0-finder1.png 400 × 400; 68 KB
- DNA Builder1.png 858 × 248; 50 KB
- DNA Builder2.png 859 × 279; 31 KB
- DNA Builder3.png 860 × 725; 39 KB
- DNA Builder4.png 858 × 724; 69 KB
- DNA Builder5.png 859 × 727; 115 KB
- DNAcartoon ringfinder0 ringmode3 laddermode1 nucleicacidmode0.png 487 × 564; 55 KB
- DNAcartoon ringfinder1 ringmode3 laddermode0 nucleicacidmode0.png 487 × 564; 130 KB
- DNAcartoon ringfinder1 ringmode3 laddermode0 nucleicacidmode1.png 487 × 564; 116 KB
- DNAcartoon ringfinder1 ringmode3 laddermode1 nucleicacidmode0.png 487 × 564; 137 KB
- DNAcartoon ringfinder2 ringmode3 laddermode1 nucleicacidmode0.png 487 × 564; 124 KB
- DNAcartoon ringfinder3 ringmode3 laddermode1 nucleicacidmode0.png 487 × 564; 137 KB
- DNAcartoon ringmode0 laddermode1 ringfinder1 nucleicacidmode0.png 487 × 564; 101 KB
- DNAcartoon ringmode1 laddermode1 ringfinder1 nucleicacidmode0.png 487 × 564; 110 KB
- DNAcartoon ringmode2 laddermode1 ringfinder1 nucleicacidmode0.png 487 × 564; 104 KB
- DNAcartoon ringmode3 laddermode1 ringfinder1 nucleicacidmode0.png 487 × 564; 134 KB
- DashColor64f.png 640 × 480; 8 KB
- DashColorMarine.png 640 × 480; 7 KB
- DashColorRed.png 640 × 480; 6 KB
- DashGap0.png 640 × 480; 3 KB
- DashGap3.png 640 × 480; 3 KB
- DashWidthEx.jpg 384 × 378; 19 KB
- Dash square ends.png 640 × 480; 79 KB
- Dashes1000.png 640 × 480; 81 KB
- Dashes3500.png 640 × 480; 73 KB
- Default.JPG 192 × 283; 7 KB
- Default sheet.JPG 261 × 421; 18 KB
- Define system tab.PNG 390 × 420; 74 KB
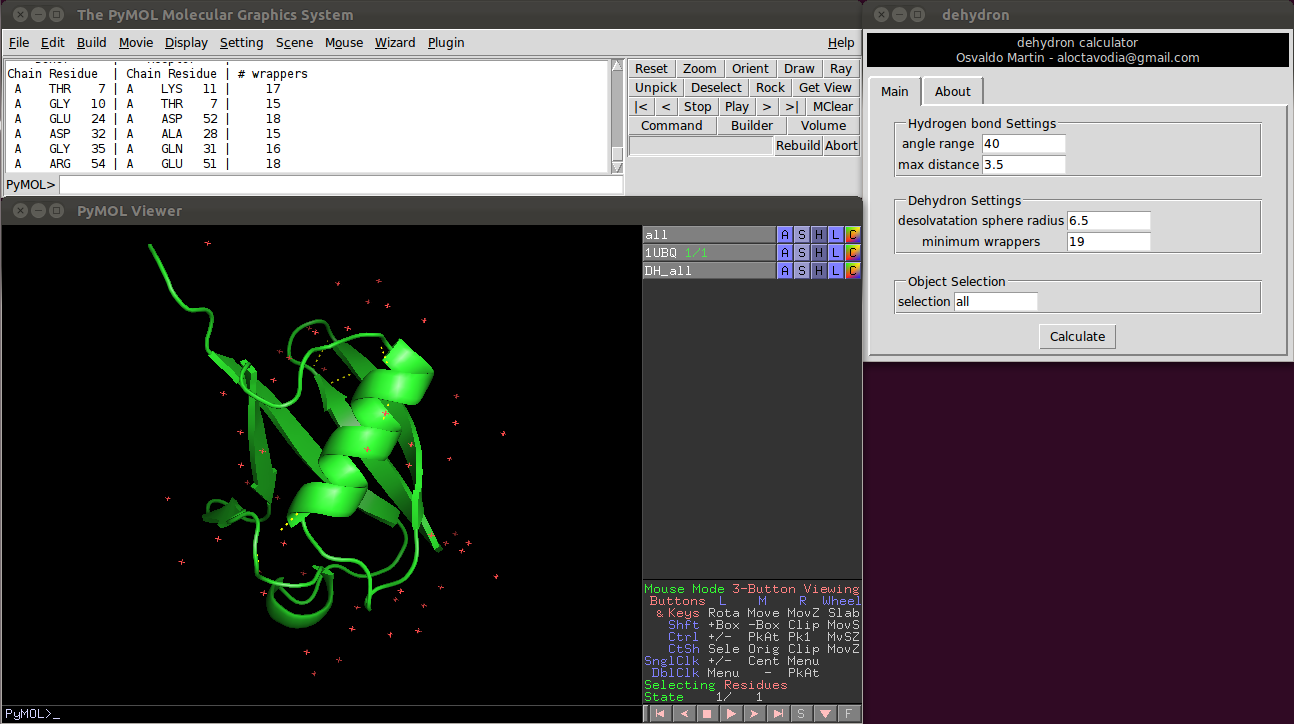
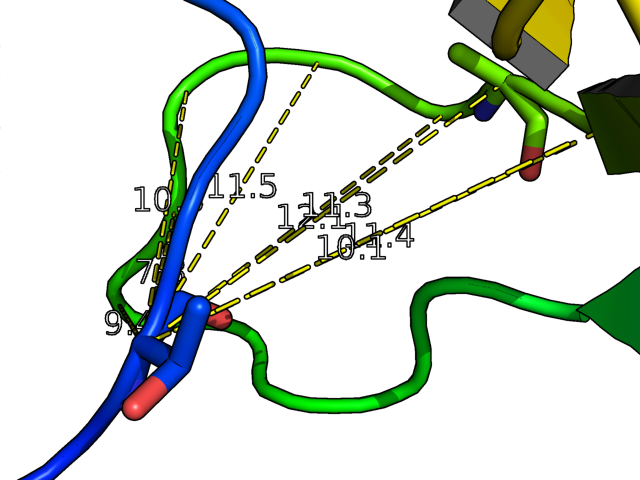
- Dehydrons.png 1,294 × 724; 125 KB
- Demo DSSP Stride plugin 1pyg.png 1,356 × 529; 364 KB
- Demo DSSP plugin 1pyg.png 1,117 × 552; 390 KB
- Density-stereo.png 1,952 × 1,000; 704 KB
- DepthCue0.png 320 × 240; 44 KB
- DepthCue1.png 320 × 240; 43 KB
- Depthmol.mpegDepthmol.mpeg File missing
- Depthmol.mpeg.zip ; 6.85 MB
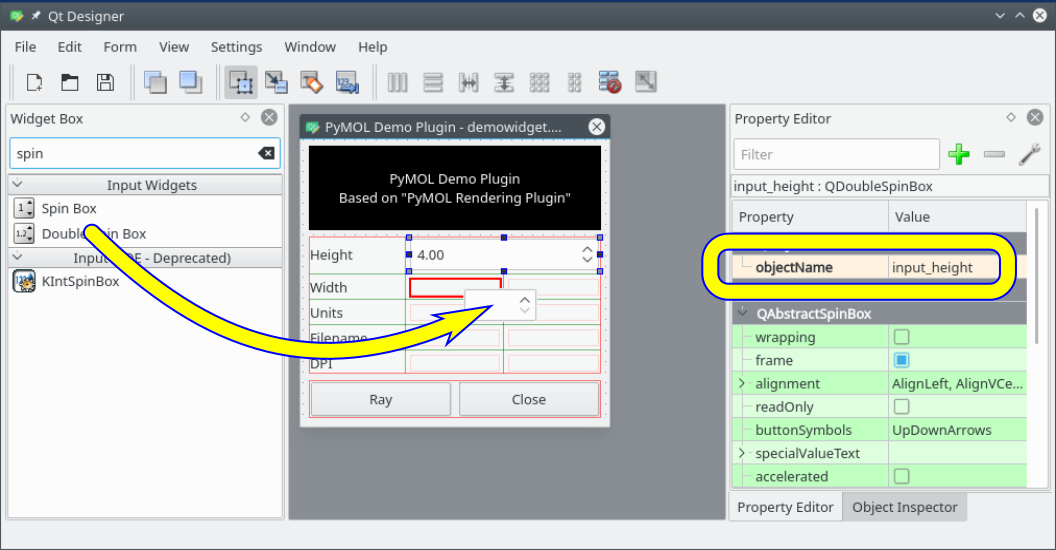
- Designer-demo-plugin.png 1,056 × 550; 146 KB
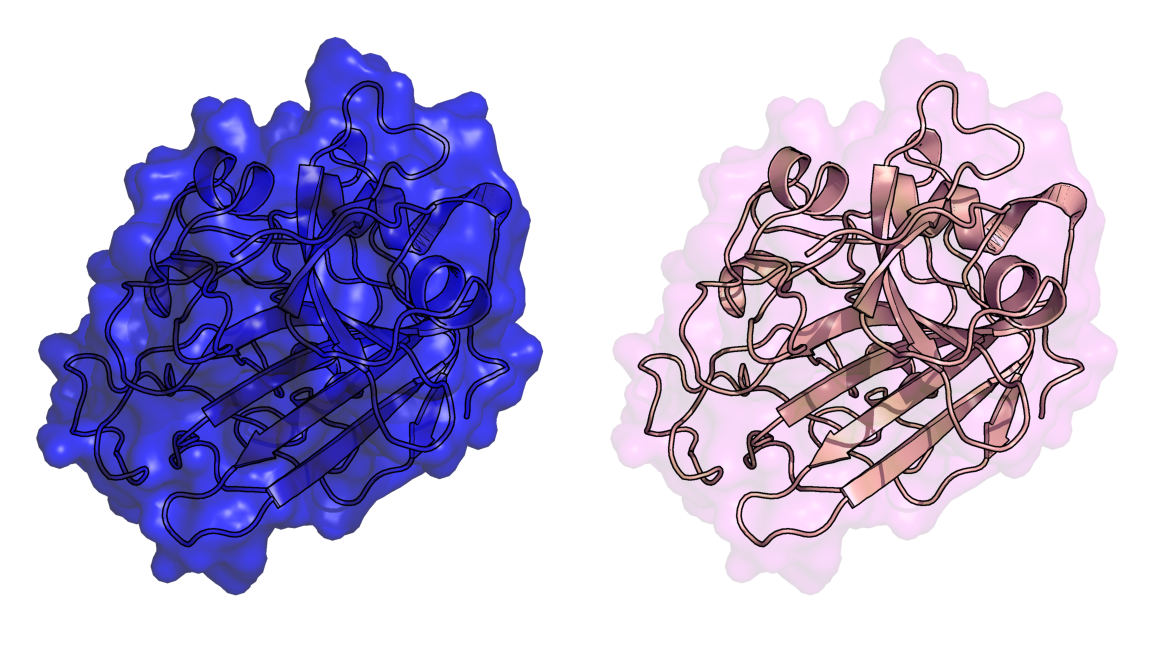
- Diffy transp.png 1,149 × 646; 644 KB
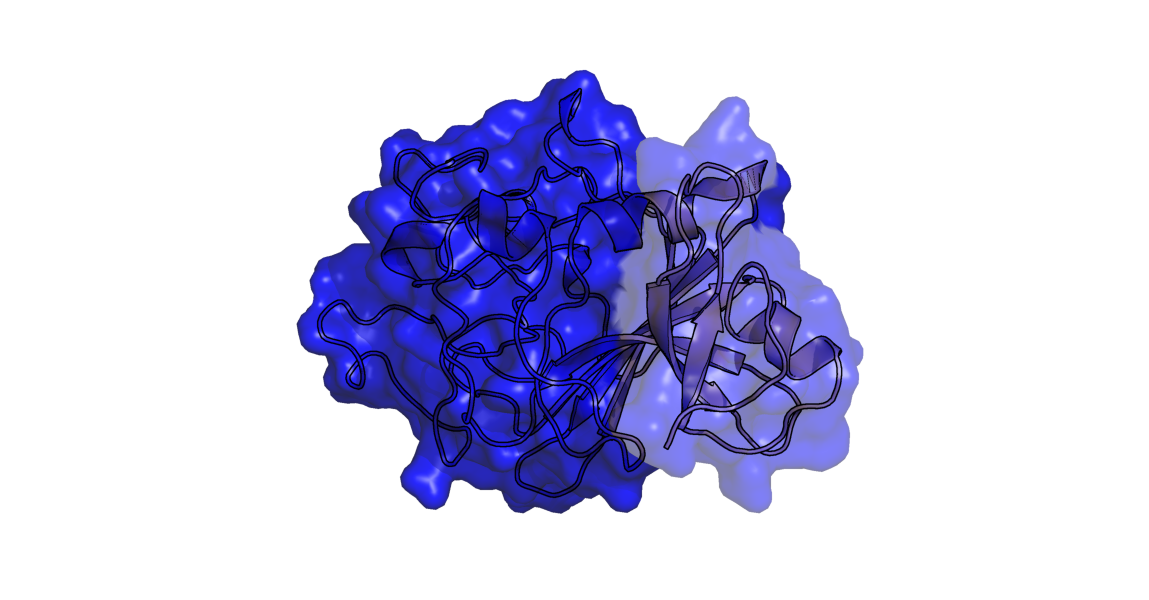
- Diffy transp2.png 1,149 × 600; 292 KB
- Dist ex1.png 988 × 728; 163 KB
- Dist ex2.png 640 × 480; 115 KB
- DistancesRH ex1.png 1,654 × 761; 271 KB
- DistancesRH ex2.png 1,770 × 601; 262 KB
- DistancesRH ex3.png 669 × 628; 190 KB
- Distancetoatom 0.png 800 × 600; 354 KB
- Distancetoatom 1.png 800 × 600; 189 KB
- Distancetoatom 2.png 800 × 600; 116 KB
- Dl3500.png 640 × 480; 74 KB
- Dots ex.png 640 × 480; 471 KB
- DoubleBonds.png 800 × 600; 29 KB
- DrawMinBB.png 800 × 600; 206 KB
- DrawMinBoundBox.tar.bz2DrawMinBoundBox.tar.bz2 File missing
- Draw ex.png 1,600 × 1,482; 1.33 MB
- Drawgridbox.png 600 × 600; 161 KB
- Drawgridbox padding.png 600 × 600; 176 KB
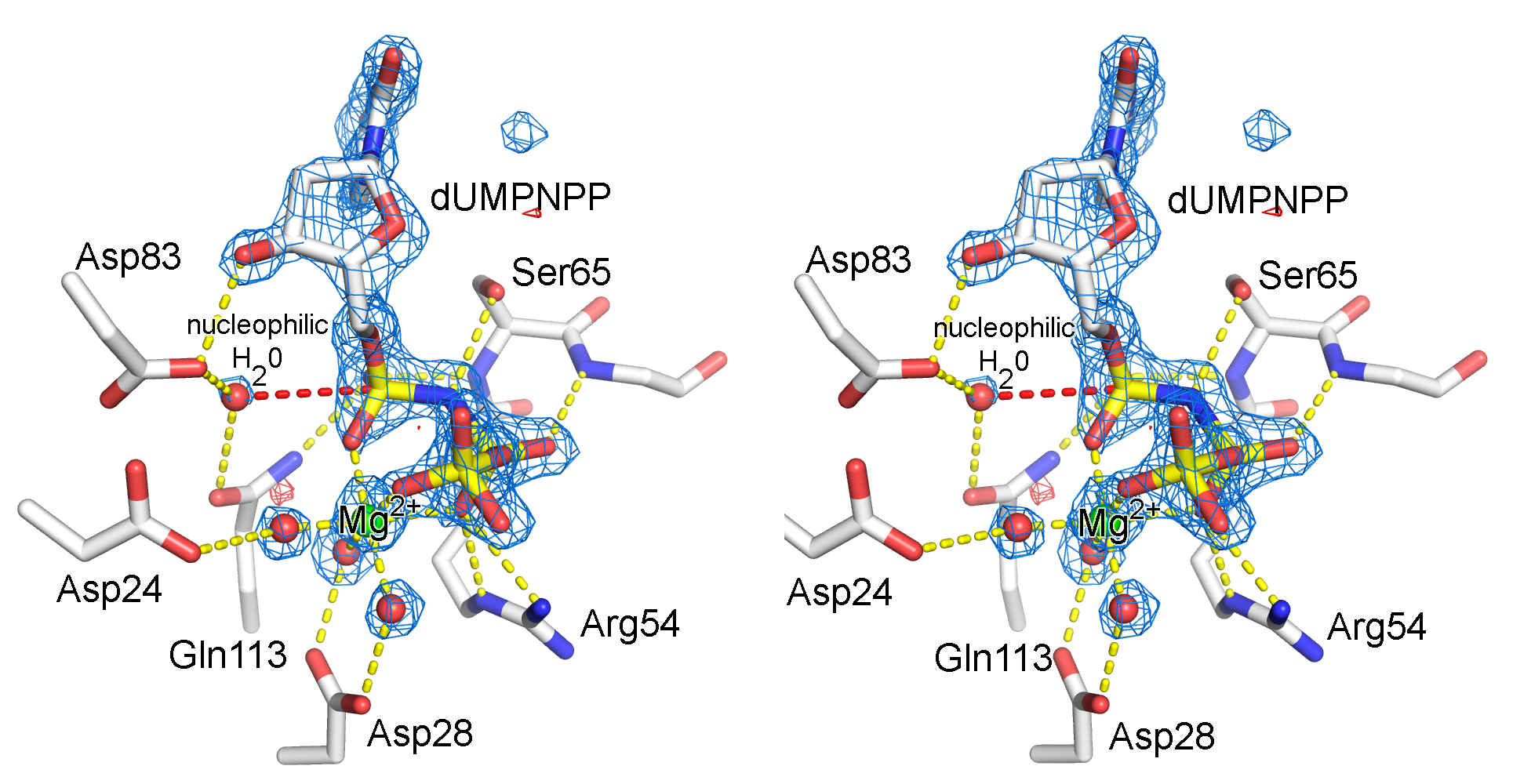
- Dutpase active02-labeled.png 706 × 820; 314 KB
- EMovie in use.jpg 640 × 436; 122 KB
- EMovie menubar.jpg 199 × 442; 33 KB
- Elbow angle 3ghe.png 600 × 800; 191 KB
- Elines.png 800 × 800; 1.14 MB
- Ell1.png 799 × 616; 359 KB
- Ell2.png 799 × 616; 350 KB
- ErmC 2ERC surface solvent OFF.png 640 × 480; 145 KB
- ErmC 2ERC surface solvent ON.png 640 × 480; 155 KB
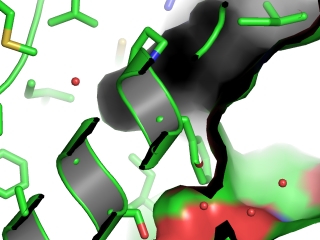
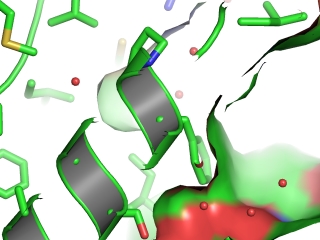
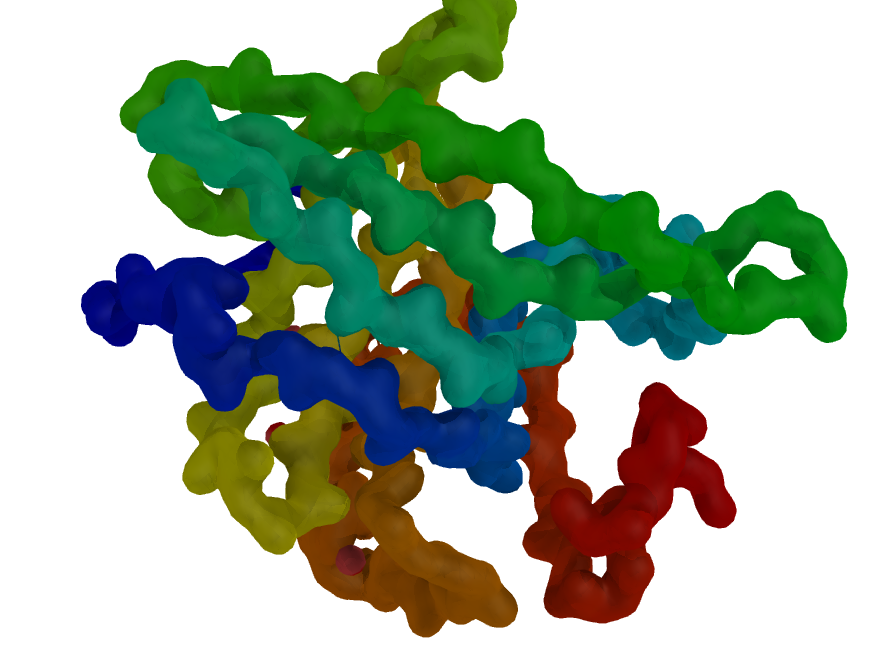
- Especially when straight features parallel to the z axis are shown, the effect may be large.
- Example.png 1,350 × 525; 470 KB
- Example 1 FAD 1C0L.png 2,000 × 1,500; 261 KB
- Extend0.png 640 × 480; 32 KB
- Extend1.png 640 × 480; 32 KB
- Extend3.png 640 × 480; 32 KB
- FEBS-585-8.png 720 × 967; 1.7 MB
- Fa.png 921 × 702; 352 KB
- Fa2.png 921 × 702; 395 KB
- Fancy.JPG 195 × 278; 7 KB
- Fancy sheet.JPG 255 × 424; 19 KB
- Fat dashes.png 640 × 480; 83 KB
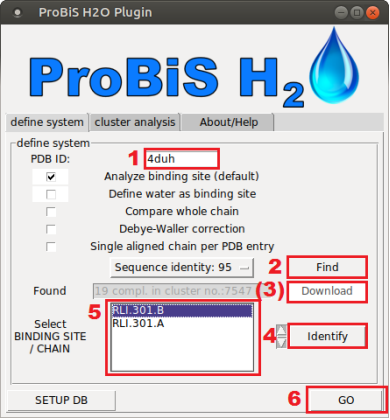
- Feature 4duh.PNG 601 × 450; 270 KB
- Figure1.png 537 × 385; 59 KB
- Figure2a.png 981 × 516; 129 KB
- File.png 300 × 223; 16 KB
- File.xyz.tar ; 10 KB
- FindExRes.png 1,304 × 831; 382 KB
- Flags-ignore-exfoliate.png 600 × 280; 64 KB
- Fnab example.png 573 × 668; 149 KB
- FocalBlur a1.0 r1.png 740 × 560; 337 KB
- FocalBlur a2.0 r1.png 740 × 560; 356 KB
- FocalBlur a4.0 r0.png 740 × 560; 380 KB
- FocalBlur a4.0 r1.png 740 × 560; 368 KB
- FocalBlur current sampling.png 400 × 400; 13 KB
- FocalBlur optimal sampling.png 400 × 400; 14 KB
- Focal blur ex6.png 640 × 480; 222 KB
- Focal blur ex ap3.png 640 × 480; 74 KB
- Focal blur ex ap3 mode1.png 640 × 480; 154 KB
- Focal blur ex ap3 mode2.png 640 × 480; 78 KB
- Focal blur ex ap3 mode3.png 640 × 480; 129 KB
- Fog off.png 640 × 480; 85 KB
- Fog on.png 640 × 480; 97 KB
- Fog start 1.png 640 × 480; 90 KB
- Fog start default.png 640 × 480; 102 KB
- Fog start minus0.5.png 640 × 480; 105 KB
- Fogoff.png 640 × 480; 131 KB
- Fogon.png 640 × 480; 135 KB
- Font ex.png 800 × 600; 80 KB
- For many images, the difference is hardly visible.For many images, the difference is hardly visible. File missing
- Fov60.png 640 × 480; 132 KB
- Fragment ex.png 640 × 480; 14 KB
- Frame slider.png 410 × 48; 3 KB
- Frizz.JPG 395 × 258; 8 KB
- GLSLToonCartoon.png 1,024 × 768; 484 KB
- GLSLToonSperes.png 1,024 × 768; 400 KB
- GLURotamerComparison5.png 598 × 388; 79 KB
- Gallery blob sc.png 1,097 × 815; 243 KB
- Gallery dark tube mc.png 875 × 653; 269 KB
- Geo measures.png 842 × 497; 8 KB
- Glgui1.png 249 × 547; 9 KB
- Glgui2.png 243 × 553; 8 KB
- Glgui3.png 241 × 559; 9 KB
- Glgui4.png 860 × 498; 8 KB
- Glgui5.png 860 × 498; 8 KB
- Glgui6.png 860 × 498; 8 KB
- Glgui7.png 860 × 498; 8 KB
- Glgui8.png 860 × 498; 11 KB
- Gm0.png 1,000 × 1,000; 444 KB
- Gm1.png 1,000 × 1,000; 385 KB
- Gm2.png 1,000 × 1,000; 111 KB
- Goldenberg cover.jpg 714 × 864; 273 KB
- Goodsell like2.png 800 × 615; 329 KB
- GridMax1.png 300 × 300; 22 KB
- GrideMax0.png 300 × 300; 25 KB
- Group off.png 257 × 169; 3 KB
- Group on1.png 240 × 96; 2 KB
- Group on2.png 238 × 126; 3 KB
- Gslike.png 800 × 800; 413 KB
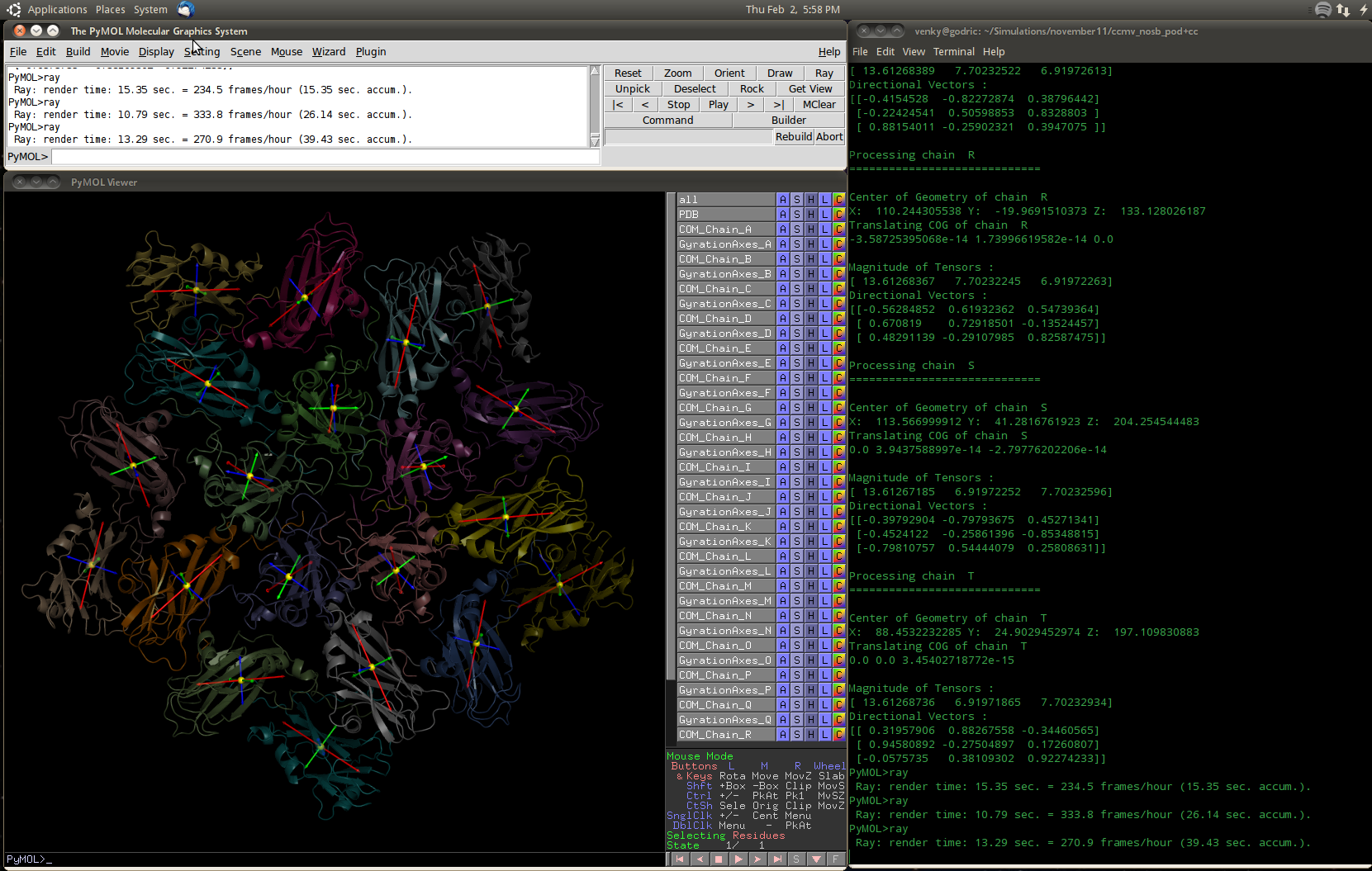
- Gyration tensor.png 1,661 × 1,055; 578 KB
- H.a.steinberg JBC 08.jpgH.a.steinberg JBC 08.jpg File missing
- H.a.steinberg jbc 08 sm.gifH.a.steinberg jbc 08 sm.gif File missing
- H2O.png 1,000 × 1,000; 88 KB
- Half-yes.png 250 × 184; 10 KB
- Halfbond-off.jpg 250 × 184; 4 KB
- Hdgyp cover.jpg 1,079 × 1,418; 580 KB
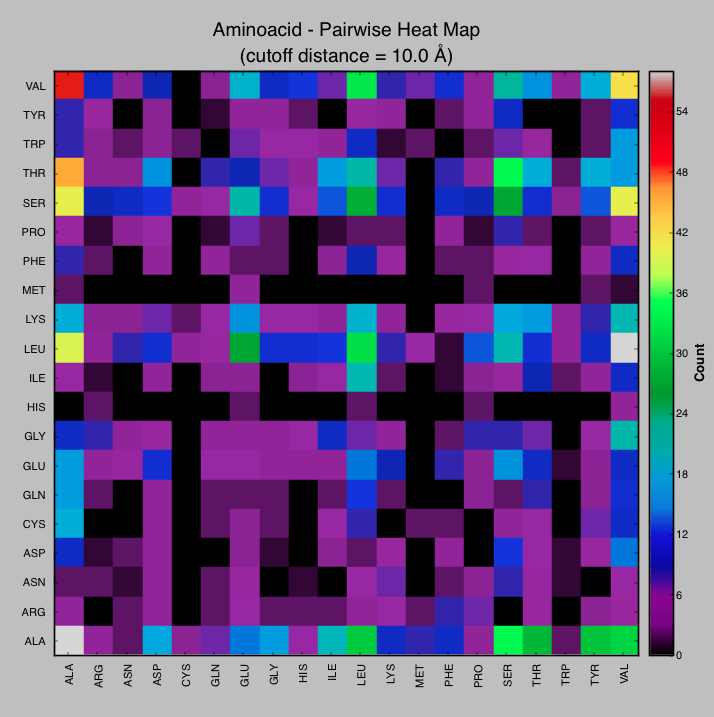
- Heatmap-CMPyMOL.PNG 700 × 691; 46 KB
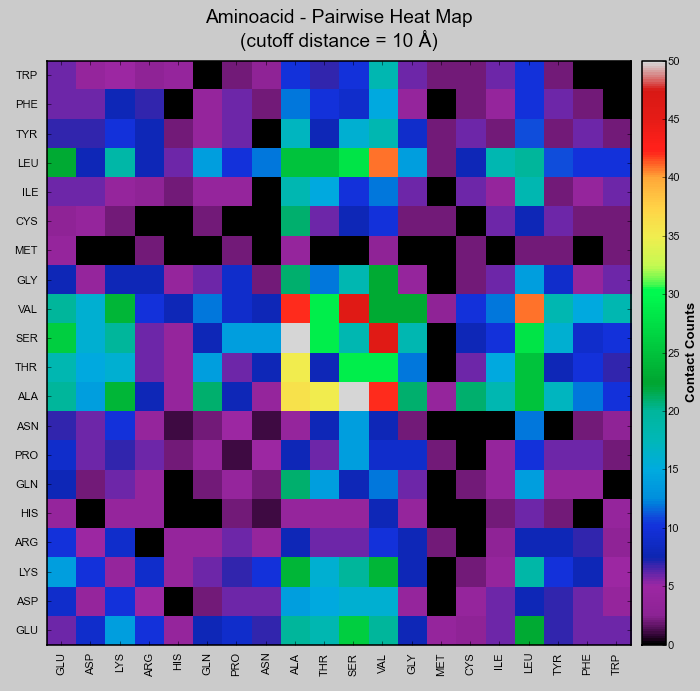
- Heatmap.png 714 × 717; 53 KB
- Helicity check-1.0.tar.bz2Helicity check-1.0.tar.bz2 File missing
- Helicity check-1.0.tar.zip ; 2 KB
- Helix orientation1.png 807 × 616; 67 KB
- Helix orientation2.png 807 × 616; 76 KB
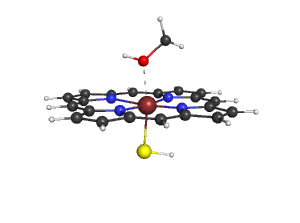
- Heme18.png 333 × 250; 143 KB
- HhaExample.png 1,000 × 750; 603 KB
- HhaI20example.png 1,600 × 1,200; 1.28 MB
- Highlight ss1.png 819 × 625; 236 KB
- Hydrobond action.png 861 × 984; 187 KB
- Iapp.gif 339 × 440; 105 KB
- Ig.png 355 × 1,012; 50 KB
- Ig mode0.png 1,195 × 969; 519 KB
- Ig mode1.png 1,195 × 969; 436 KB
- Ig off.png 975 × 969; 381 KB
- Ig on.png 1,195 × 969; 434 KB
- Igcs df30.png 304 × 753; 30 KB
- Igwidth220.png 1,195 × 969; 519 KB
- Igwidth50.png 1,025 × 969; 456 KB
- Igwidth500.png 1,475 × 969; 526 KB
- Image large.png 613 × 768; 335 KB
- Image merged.png 640 × 480; 260 KB
- Ind0.png 640 × 480; 81 KB
- Ind1.png 640 × 480; 58 KB
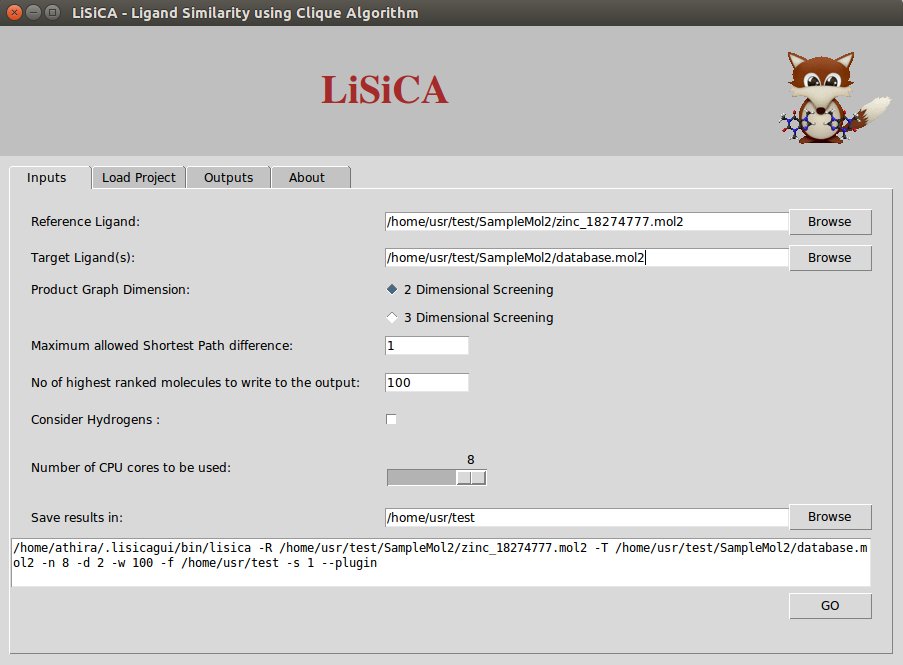
- InputTab2DLiSiCALinuxv1.0.0.png 903 × 665; 66 KB
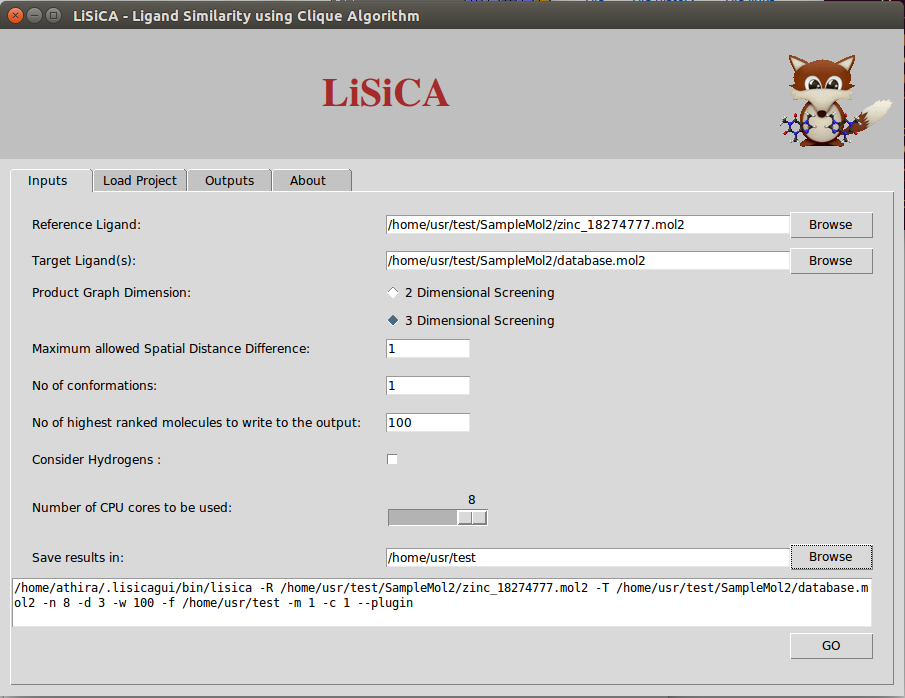
- InputTab3DLiSiCALinuxv1.0.0.png 905 × 698; 70 KB
- InteranlGuiControlSize Default.png 249 × 438; 10 KB
- Interface02 labeled.png 713 × 713; 338 KB
- InternalFeedback1.png 1,195 × 861; 21 KB
- InternalFeedback10.png 1,195 × 969; 520 KB
- Intra xfit-1nmr.png 600 × 600; 174 KB
- Irvinggeis pymol 500px.png 500 × 454; 381 KB
- Isomesh-buffer-carve.png 281 × 289; 25 KB
- Isomesh ex1.png 1,120 × 882; 1.76 MB
- It.png 765 × 592; 154 KB
- JBC 080406 no31 proof.png 1,650 × 2,175; 3.21 MB
- JBC Apr18 2008.jpg 472 × 622; 306 KB
- JExpBiol-214-4.jpg 600 × 776; 233 KB
- J square.jpg 432 × 458; 176 KB
- Jbc 20070413.gif 117 × 150; 10 KB
- Jbc mchu.jpeg 623 × 811; 102 KB
- Jctcce v011i009.jpg 2,456 × 3,262; 1.73 MB
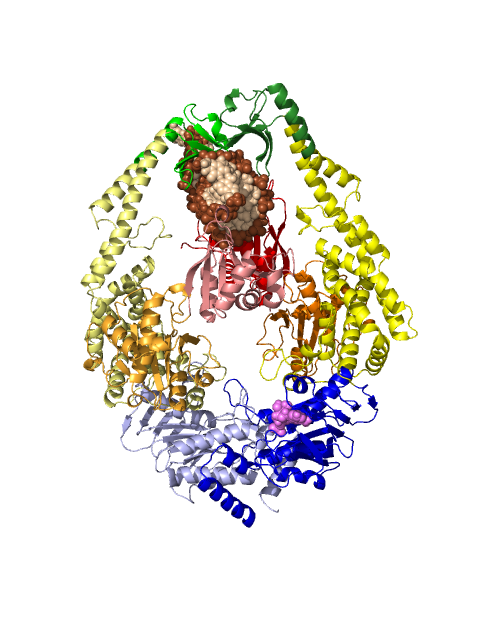
- Kchannel-rainbow.png 457 × 512; 146 KB
- KscA.png 218 × 300; 83 KB
- L1.png 965 × 815; 320 KB
- L2.png 965 × 815; 318 KB
- Label ex.png 1,052 × 718; 324 KB
- Label pre.png 1,052 × 718; 122 KB
- Large surface.png 600 × 800; 423 KB
- Large surface approx.png 600 × 800; 221 KB
- Largecover.gif 440 × 580; 105 KB
- Lce.png 496 × 200; 11 KB
- Libgfortran.3.dylib.bz2Libgfortran.3.dylib.bz2 File missing
- Libgfortran.3.dylib.zip ; 264 KB
- Light count 0.png 640 × 480; 138 KB
- Light count 10.png 640 × 480; 154 KB
- Light count 2.png 640 × 480; 146 KB
- Lines ex.png 640 × 480; 140 KB
- Ll.gif 320 × 200; 310 KB
- LoadBfacts02.png 1,042 × 877; 248 KB
- LoadBfacts1.png 1,071 × 858; 336 KB
- Load brick example.png 400 × 400; 45 KB
- Long movie ray.mpg ; 5.98 MB
- Ls0.png 873 × 459; 84 KB
- Ls2.png 873 × 459; 88 KB
- Lw1.png 930 × 679; 310 KB
- Lw2.png 930 × 679; 337 KB
- MMcoverMay2009.gif 95 × 125; 10 KB
- MMcoverMay2009.png 95 × 125; 12 KB
- Map double.png 800 × 654; 783 KB
- Map double2.png 800 × 654; 947 KB
- Map double3.png 800 × 654; 966 KB
- Map normal.png 800 × 654; 582 KB
- Map set ex.png 934 × 707; 229 KB
- Mapping example1.png 333 × 250; 112 KB
- Measure.png 1,147 × 760; 264 KB
- Menu.png 775 × 561; 292 KB
- Menu1.png 1,070 × 476; 129 KB
- Mesh ex.png 400 × 400; 325 KB
- Mesh quality 0.jpg 640 × 480; 250 KB
- Mesh quality 1.jpg 640 × 480; 224 KB
- Mesh rm0.png 1,066 × 787; 1.16 MB
- Mesh rm3.png 1,066 × 787; 471 KB
- Mesh type 1.png 640 × 480; 70 KB
- Mesh type 1 ray.jpg 640 × 480; 143 KB
- Mesh w05.png 1,066 × 787; 1.15 MB
- Mesh width 3.jpg 640 × 480; 157 KB
- Middle ray.png 300 × 300; 58 KB
- Min mesh spacing 2.jpg 640 × 480; 155 KB
- Min mesh spacing default.jpg 640 × 480; 222 KB
- Modevector.png 1,600 × 673; 272 KB
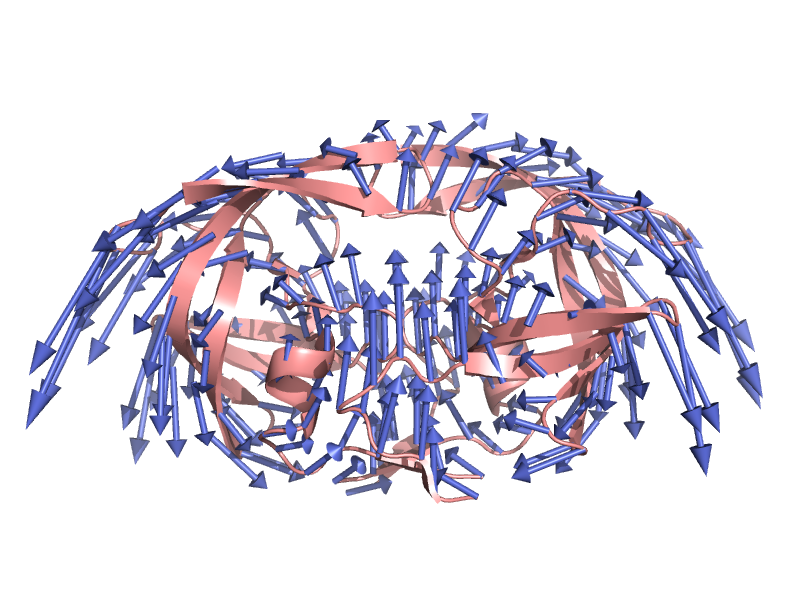
- Modevectors.png 1,380 × 1,162; 487 KB
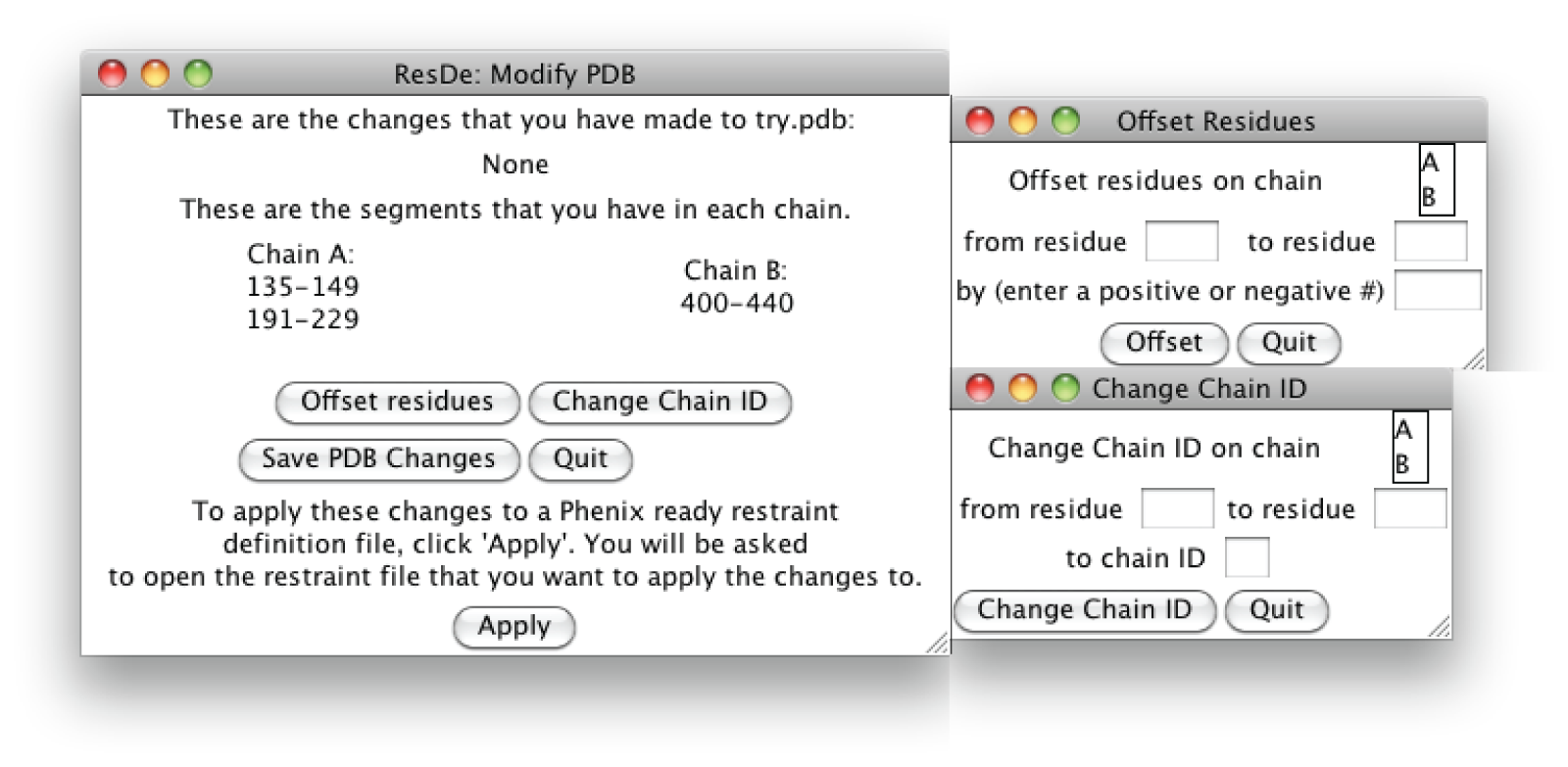
- ModifyPDB all.png 1,600 × 783; 221 KB
- MolecularCell011.jpg 2,413 × 3,255; 3.02 MB
- Molelogo.png 125 × 125; 8 KB
- Motions movie.mpg ; 1.88 MB
- Mouse-3view.png 222 × 102; 2 KB
- Movie.gif 700 × 500; 2.11 MB
- Movie color fade example 1.gif 300 × 200; 541 KB
- Movie fade example.gif 300 × 200; 1.29 MB
- Movies.png 595 × 152; 7 KB
- MtsslDock Import Labels.jpg 847 × 746; 91 KB
- MtsslDock distanceTable.jpg 846 × 746; 80 KB
- MtsslDock results.jpg 996 × 1,034; 265 KB
- MtsslDock screenshot.jpg 1,711 × 1,259; 329 KB
- MtsslTrilaterate screenshot.png 605 × 792; 317 KB
- MtsslWizard gui.jpg 815 × 445; 303 KB
- MtsslWizardv1-1.jpg 1,051 × 1,087; 282 KB
- Multilayer.png 640 × 480; 279 KB
- MuraEtAl PLoS BiomolGraphics SuppFigS3 01nov2011 erratum.pdf 0 × 0; 7.38 MB
- Mutag1.png 1,280 × 1,024; 127 KB
- Mutag2.png 1,280 × 1,024; 130 KB
- Mutag3.png 1,280 × 1,024; 130 KB
- Mutag4.png 1,280 × 1,024; 117 KB
- Mutagsmall3.png 600 × 480; 124 KB
- Mv default.png 640 × 480; 110 KB
- Mv fig10.png 640 × 480; 102 KB
- Mv fig11.png 640 × 480; 117 KB
- Mv fig12.png 640 × 480; 101 KB
- Mv fig13.png 640 × 480; 132 KB
- Mv fig2.png 640 × 480; 131 KB
- Mv fig3.png 640 × 480; 116 KB
- Mv fig4.png 640 × 480; 108 KB
- Mv fig5.png 640 × 480; 116 KB
- Mv fig6.png 640 × 480; 109 KB
- Mv fig7.png 640 × 480; 111 KB
- Mv fig8.png 640 × 480; 97 KB
- Mv fig9.png 640 × 480; 99 KB
- Nb spheres.png 600 × 600; 186 KB
- New fonts.jpeg 640 × 480; 148 KB
- No cls.png 870 × 526; 31 KB
- No fov.png 640 × 480; 132 KB
- No persp.png 200 × 200; 10 KB
- No ray trace.png 640 × 480; 321 KB
- No shade.png 937 × 732; 363 KB
- No shader.jpg 400 × 300; 29 KB
- Noray.png 640 × 480; 181 KB
- Notcylindre.JPG 164 × 203; 4 KB
- Nucl. Acids Res.-2015-Front-Matter DBissue.jpg 1,439 × 1,765; 401 KB
- Nucleic1.png 320 × 480; 128 KB
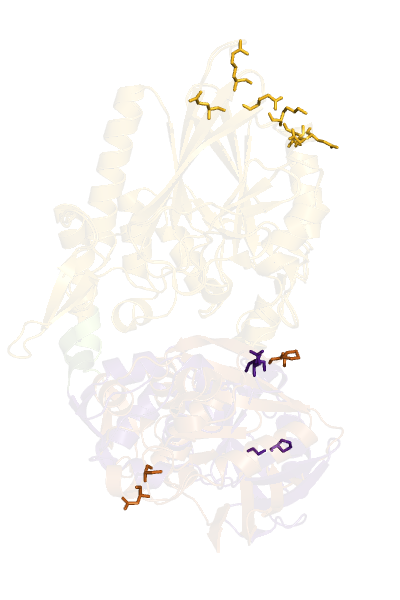
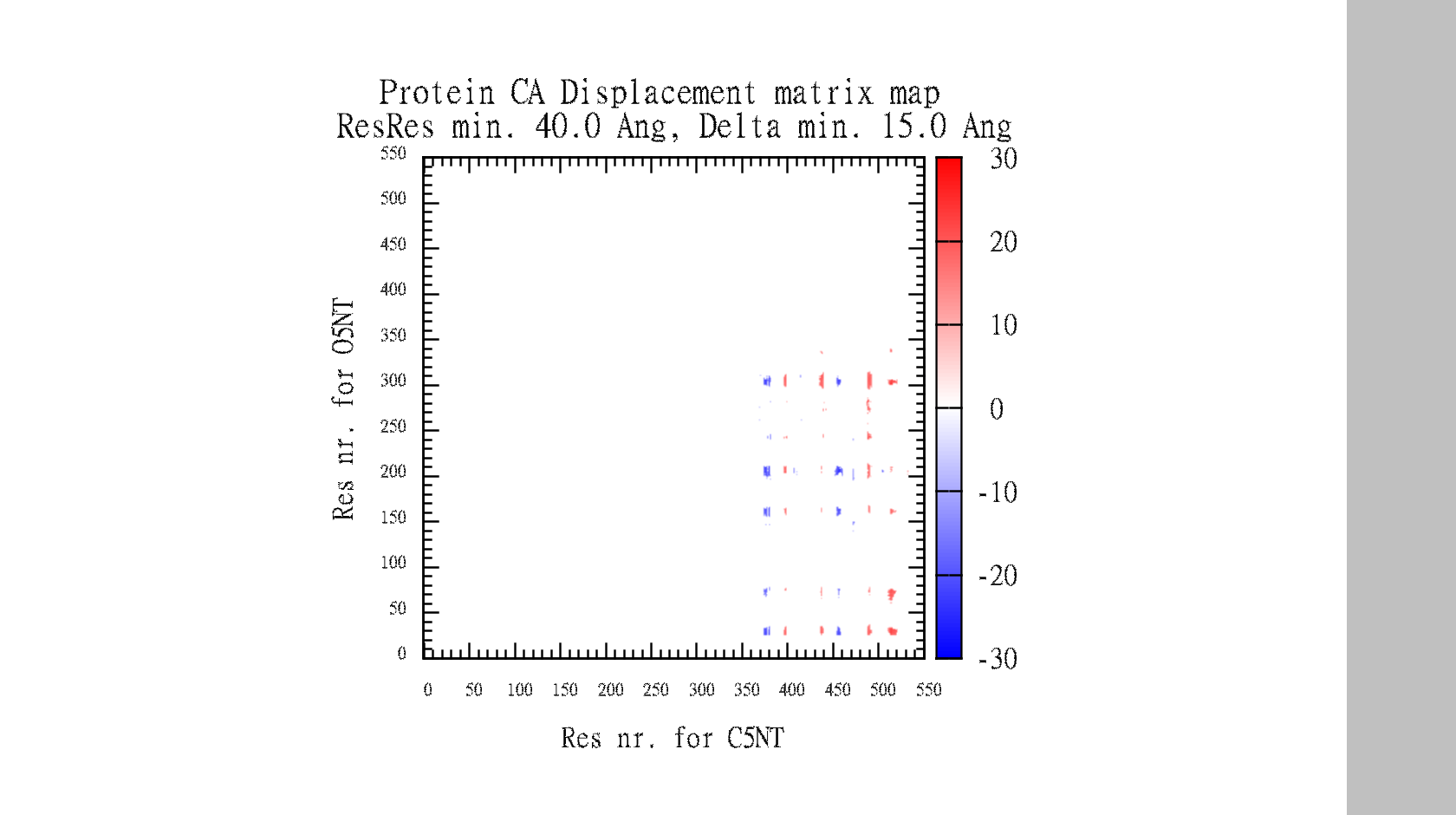
- O5NT-1HP1-A-C5NT-1HPU-C-CA-dist-map.png 640 × 387; 11 KB
- O5NT-1HP1-A-C5NT-1HPU-C-CA-dist.png 400 × 600; 106 KB
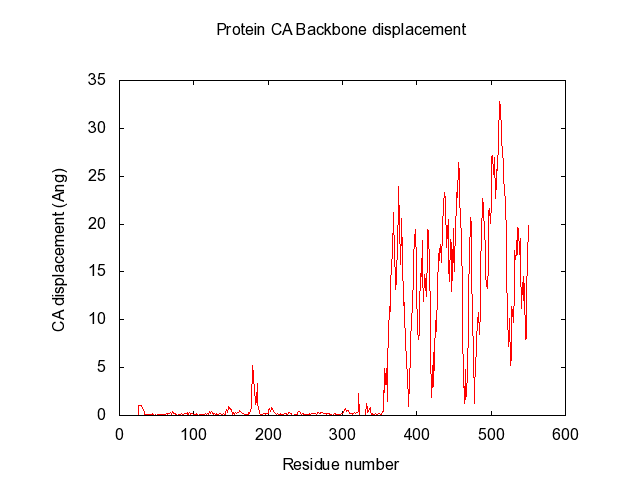
- O5NT-C5NT-CA-dist-backbone.png 640 × 480; 7 KB
- O5NT-C5NT-CA-dist-gnuplot.png 1,680 × 940; 70 KB
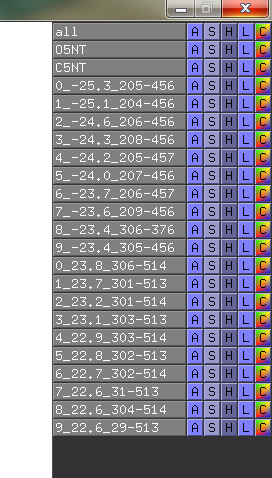
- O5NT-C5NT-CA-dist-menu.png 272 × 478; 14 KB
- O5NT-C5NT-CA-dist.png 1,460 × 970; 328 KB
- OptAlign1.png 400 × 400; 116 KB
- OptAlign2.png 400 × 400; 40 KB
- Optimize.png 862 × 721; 163 KB
- Orthooff.png 322 × 298; 55 KB
- Orthooff2.png 322 × 298; 66 KB
- Orthoon.png 322 × 298; 49 KB
- Orthoon2.png 322 × 298; 66 KB
- Orthoscopic close on off2.png 642 × 264; 98 KB
- Orthoscopic off.png 642 × 527; 136 KB
- Orthoscopic on.png 642 × 527; 139 KB
- Orthoscopic on off2.png 400 × 264; 102 KB
- Out.png 800 × 600; 199 KB
- Outline base.png 640 × 480; 166 KB
- Outline cleaned.png 640 × 480; 168 KB
- Outline composite.png 640 × 480; 170 KB
- Outline overlay.png 640 × 480; 7 KB
- Overlay03.png 1,071 × 1,051; 434 KB
- P4p6 small.jpg 353 × 353; 55 KB
- PDB 5j96 assemblies.png 1,914 × 1,054; 1.43 MB
- PDB plugin.png 856 × 230; 51 KB
- PDBe 3b43 domains 2.png 800 × 426; 79 KB
- PDF.png 640 × 478; 97 KB
- PHE delocalized.png 591 × 591; 48 KB
- PHE valence 0.png 591 × 591; 40 KB
- PHE valence 1 mode 0.png 591 × 591; 45 KB
- PHE valence 1 mode 1.png 591 × 591; 45 KB
- PICv.png 525 × 405; 55 KB
- PICv home screen.png 957 × 508; 62 KB
- PLoS BiomolGraph 2010 InputFiles.tar.bz2PLoS BiomolGraph 2010 InputFiles.tar.bz2 File missing
- PLoS BiomolGraph 2010 InputFiles.tar.zip ; 3.32 MB
- PNASCover.gif 134 × 178; 12 KB
- Pairwise1.png 1,064 × 669; 261 KB
- Pairwise2.png 1,064 × 669; 397 KB
- Pairwise3.png 1,064 × 669; 204 KB
- ParM.gif 346 × 440; 82 KB
- Patom.png 640 × 480; 148 KB
- Pdbe 2gc2 validation.png 1,366 × 727; 264 KB
- Pdbe 3b43 domains.png 1,363 × 728; 55 KB
- Pdbe 3l2p entity.png 1,366 × 725; 199 KB
- Pdbe 3unb all annotation.png 1,366 × 727; 592 KB
- Pdbe logo.png 98 × 97; 6 KB
- Persp1.png 200 × 200; 10 KB
- Persp2.png 200 × 200; 10 KB
- Phe trna small.jpg 353 × 353; 72 KB
- Pincer Angle.png 640 × 478; 52 KB
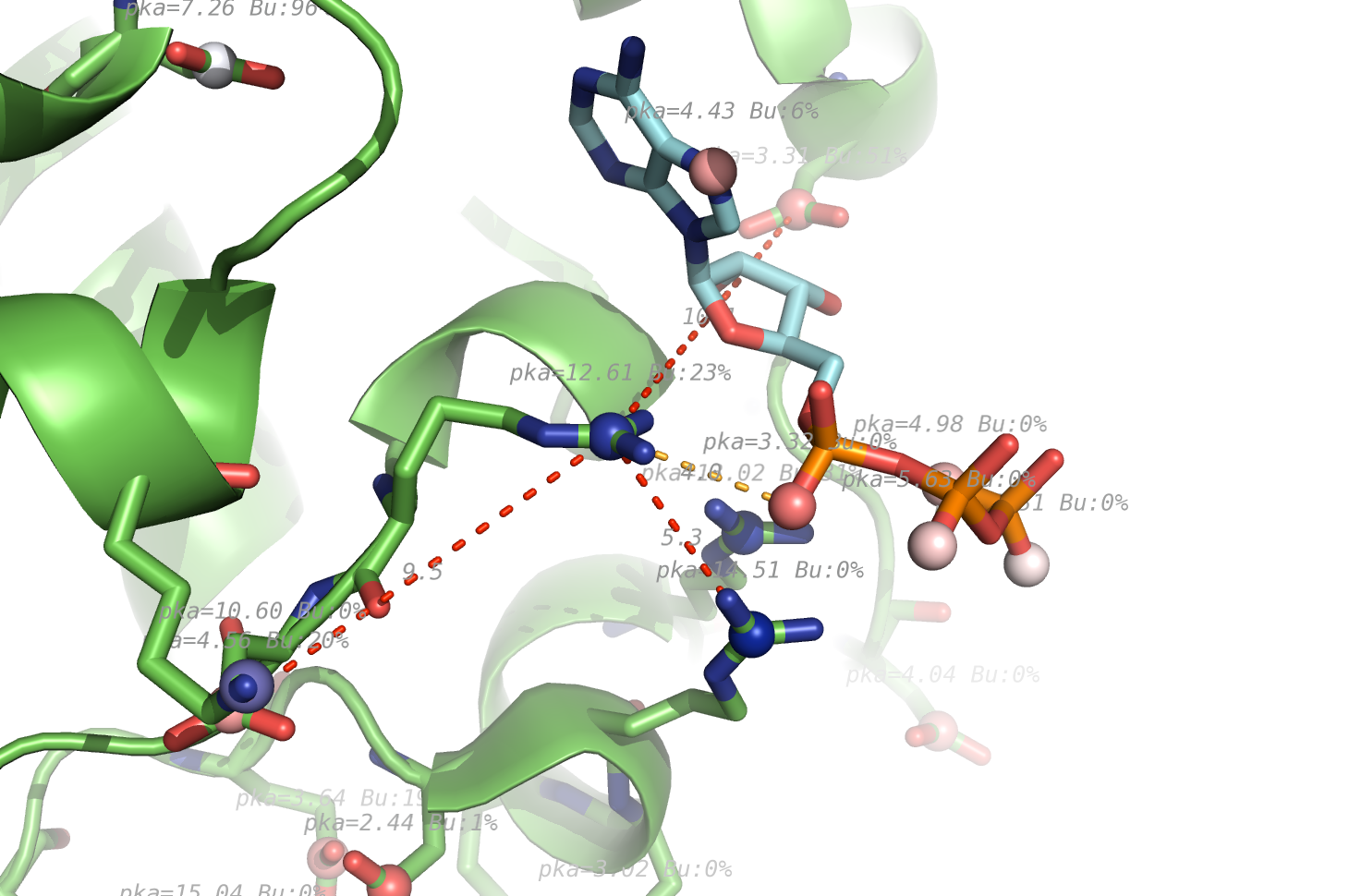
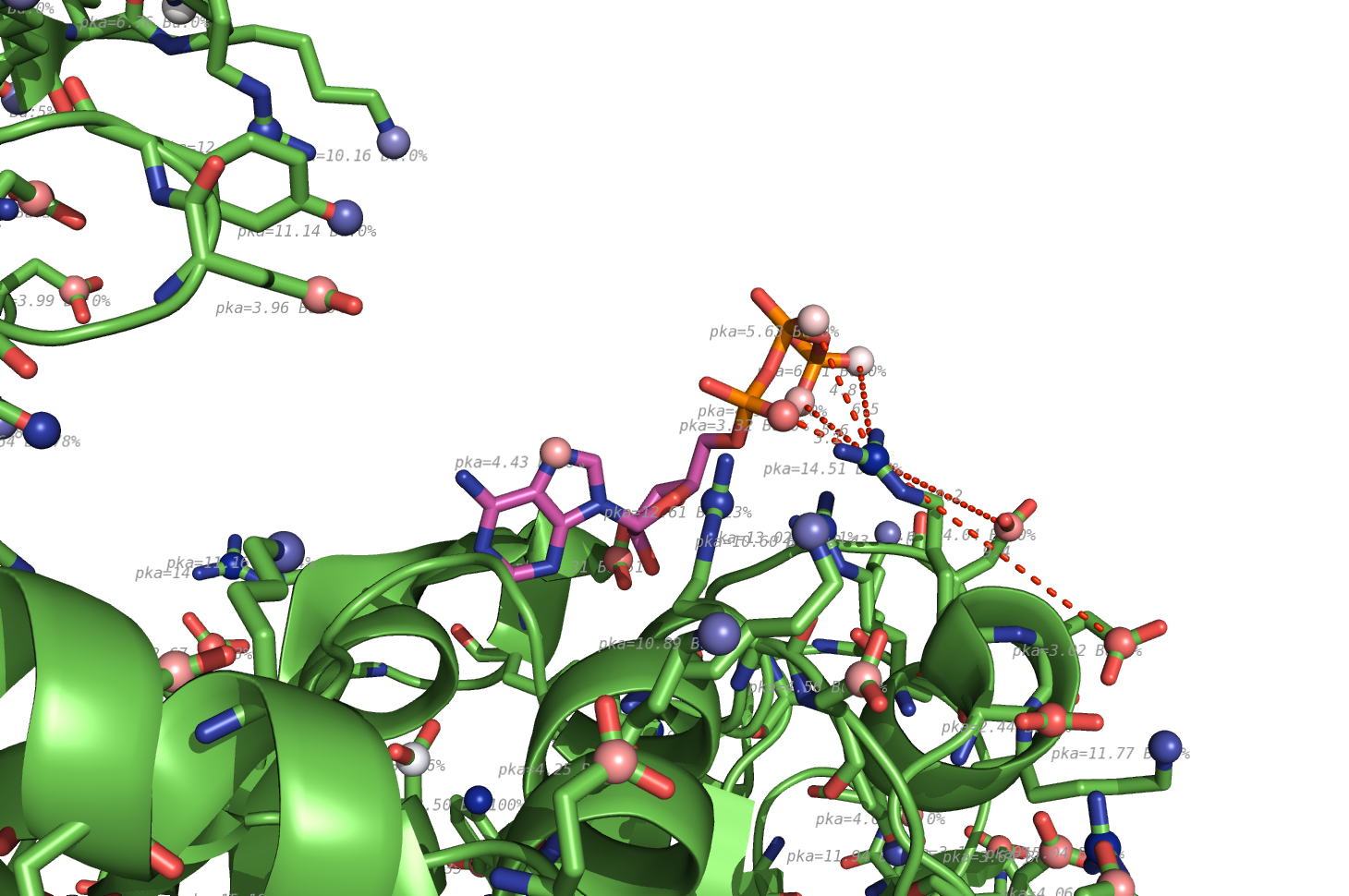
- Pkavalues.png 1,460 × 970; 587 KB
- Pl.png 873 × 459; 205 KB
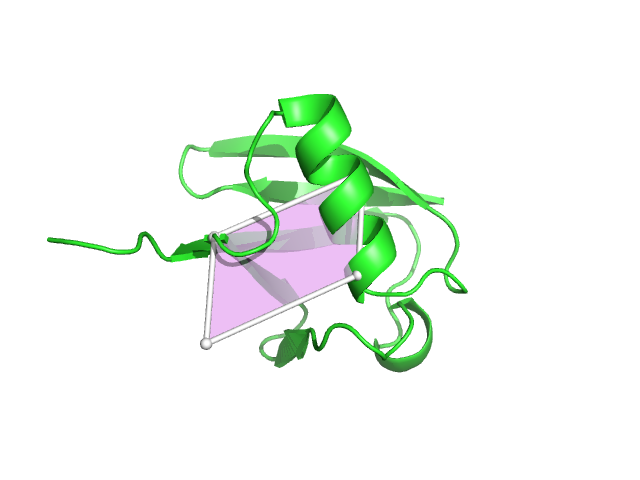
- Plane img1.png 640 × 480; 68 KB
- Plane img2.png 640 × 480; 53 KB
- Plane img3.png 640 × 480; 60 KB
- Plugin.png 1,400 × 623; 38 KB
- PluginManagerCaverPlugin301 fixed.png 670 × 478; 106 KB
- Plugin manager install new.png 461 × 393; 63 KB
- Plugin manager installed.png 461 × 393; 50 KB
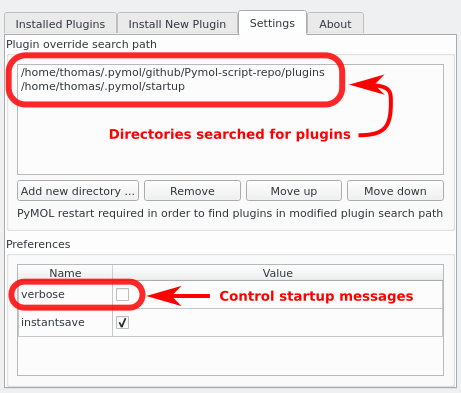
- Plugin manager settings.png 461 × 393; 38 KB
- Pocket.png 640 × 480; 668 KB
- Polar contacts small.png 800 × 600; 147 KB
- Polymer physics cover.png 594 × 792; 1,015 KB
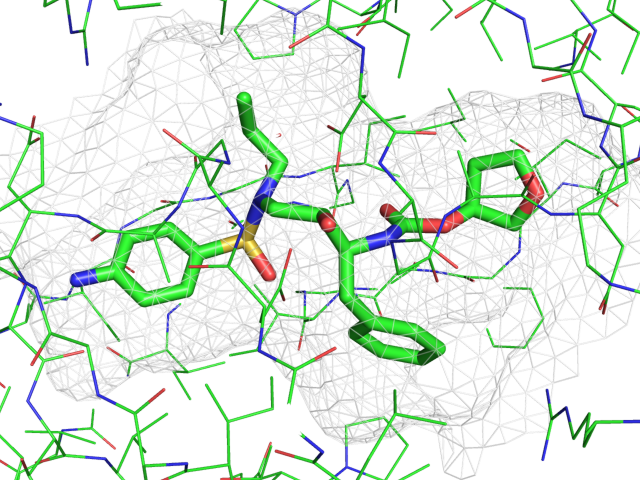
- Poseviewscreenshot.png 646 × 391; 54 KB
- Povray raytracer.png 480 × 640; 153 KB
- Properties-Inspector.png 935 × 726; 162 KB
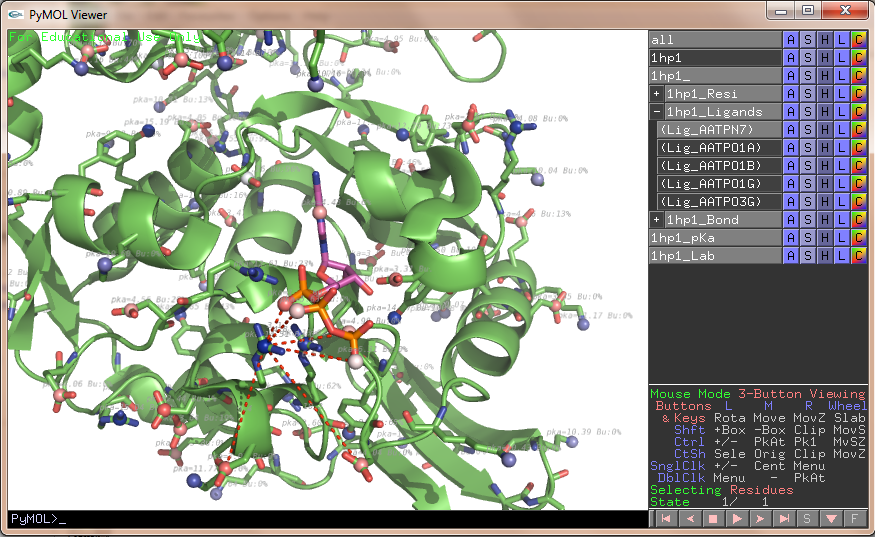
- Propka1HP1-ATP.png 1,460 × 970; 746 KB
- Propka1HP1-ZN.png 1,460 × 970; 956 KB
- Propka1HP1.png 1,460 × 970; 557 KB
- Propka1hp1.pngPropka1hp1.png File missing
- Propka1hp12.png 1,460 × 970; 573 KB
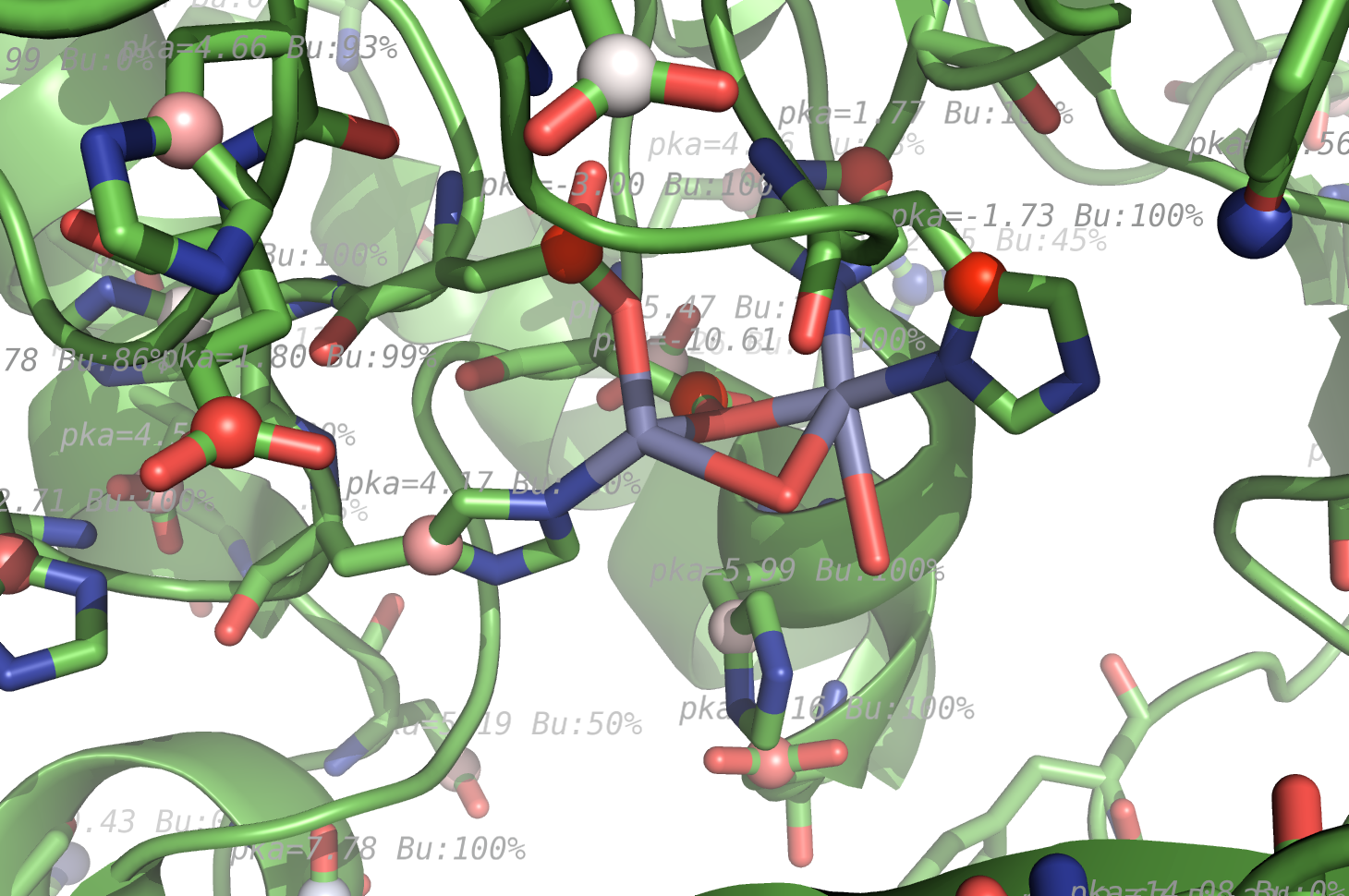
- Propka1hp13zoom2.png 1,460 × 970; 861 KB
- Propka4INS.png 1,460 × 970; 656 KB
- Propka4ins.pngPropka4ins.png File missing
- Propka4ins2.png 1,460 × 970; 626 KB
- Propkamenu3.png 773 × 560; 291 KB
- Prot contact APBS.png 640 × 480; 174 KB
- Prot contact pot.png 640 × 480; 180 KB
- Proteins 08010 c1 sp Ob.png 465 × 624; 264 KB
- Pseu1.png 1,086 × 793; 86 KB
- PyANM interface.png 599 × 409; 15 KB
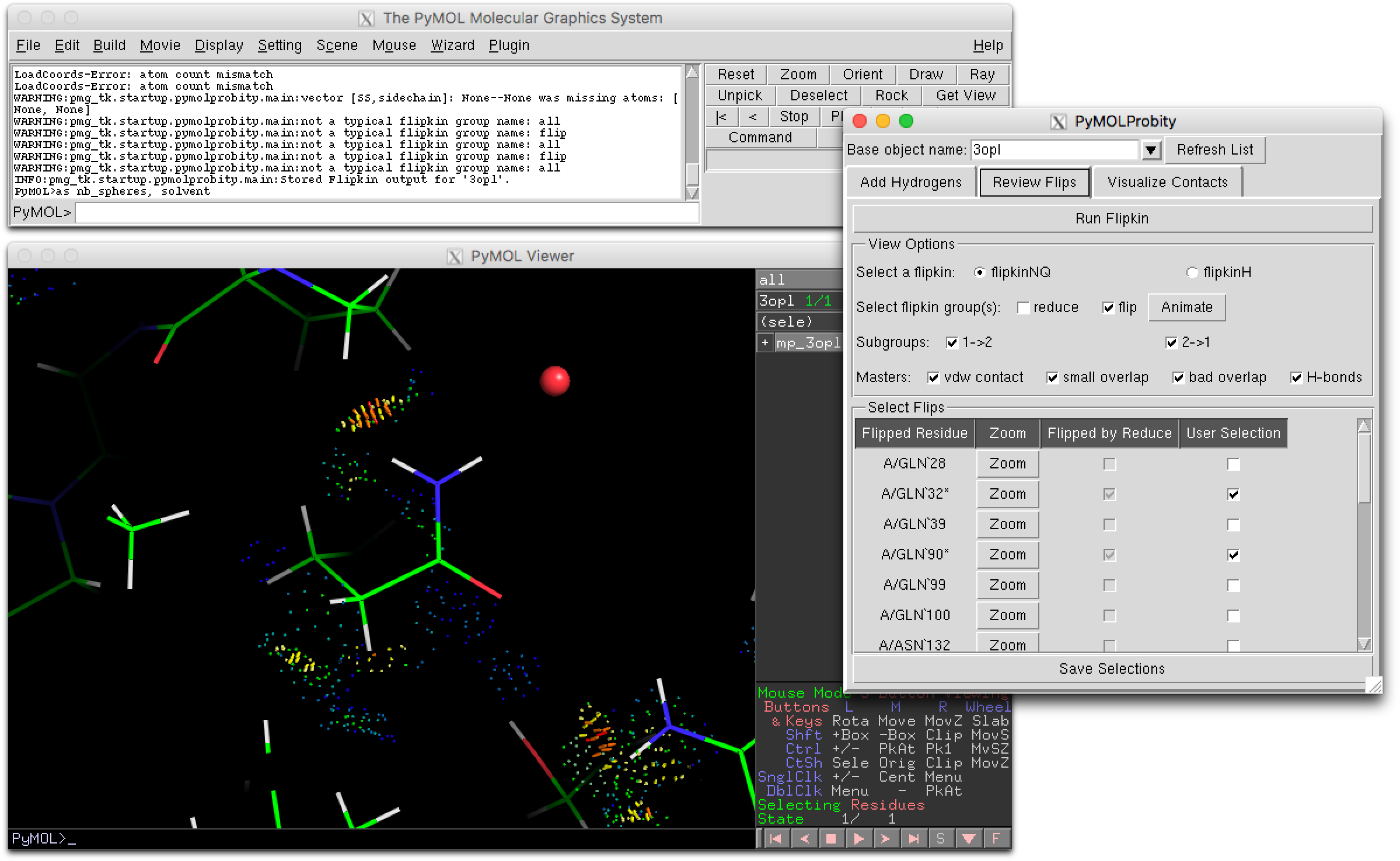
- PyMOLProbity GUI.png 2,399 × 1,476; 211 KB
- PyMOLreferenceCard-test.zip ; 68 KB
- PyMOLreferenceCard.zip ; 112 KB
- Pyanm example of arrows.png 800 × 600; 315 KB
- Pyanm example of msf.png 800 × 600; 189 KB
- Pyanm example of springs.png 800 × 600; 539 KB
- Pymol-blue magenta.png 1,058 × 100; 3 KB
- Pymol-blue red.png 1,058 × 100; 4 KB
- Pymol-blue white green.png 1,058 × 100; 4 KB
- Pymol-blue white magenta.png 1,058 × 100; 4 KB
- Pymol-rainbow.png 1,058 × 100; 10 KB
- Pymol-red blue.png 1,058 × 100; 4 KB
- Pymol.pdf 0 × 0; 168 KB
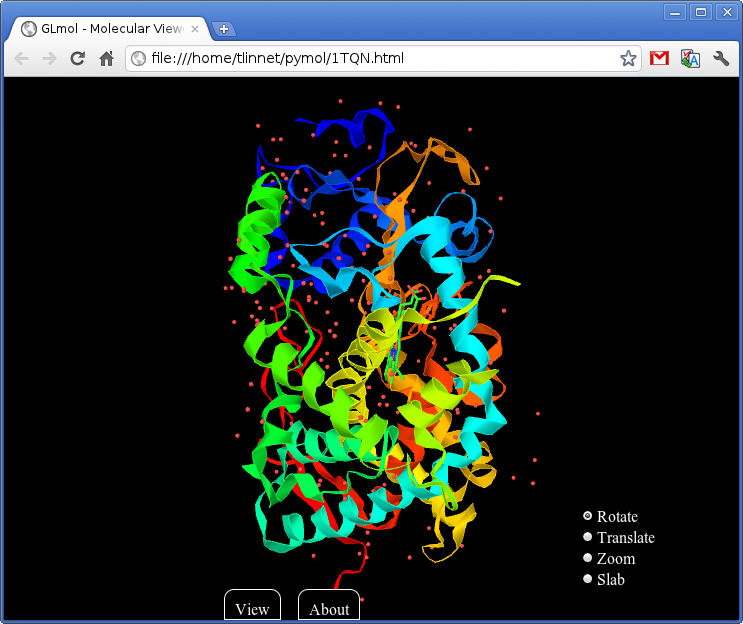
- Pymol2glmol.png 743 × 624; 142 KB
- PymolRef.pdf 0 × 0; 112 KB
- Pymol align.png 800 × 800; 321 KB
- Pymol raytracer.png 480 × 640; 178 KB
- Pymolpropkamenu.png 875 × 537; 378 KB
- Pymolwiki.png 150 × 150; 10 KB
- Pytms.gif 300 × 360; 329 KB
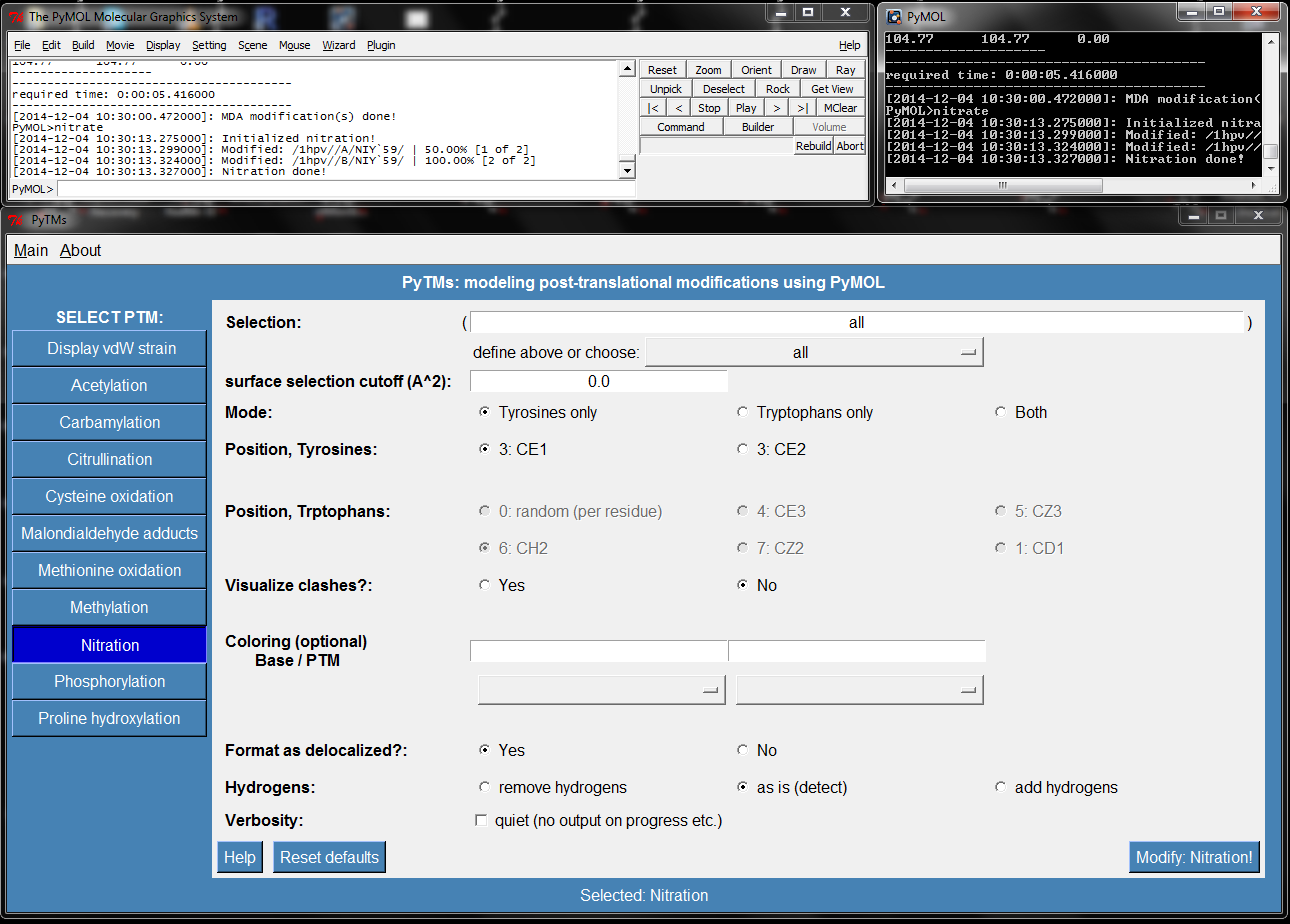
- Pytms menu.png 1,290 × 924; 154 KB
- Pytms optimization animation.gif 400 × 300; 423 KB
- Quickdisplays combo.png 472 × 295; 114 KB
- Quickdisplays disp ball stick.png 472 × 295; 95 KB
- Quickdisplays disp mesh.png 472 × 295; 117 KB
- Quickdisplays disp putty.png 472 × 295; 47 KB
- Quickdisplays disp ss.png 472 × 295; 44 KB
- Quickdisplays disp surf.png 472 × 295; 70 KB
- QuteMolLike.png 977 × 723; 437 KB
- R Cighetti AngewChemCover.gif 1,232 × 1,704; 602 KB
- Radius.png 640 × 480; 3 KB
- Radius of gyration Example.png 479 × 464; 117 KB
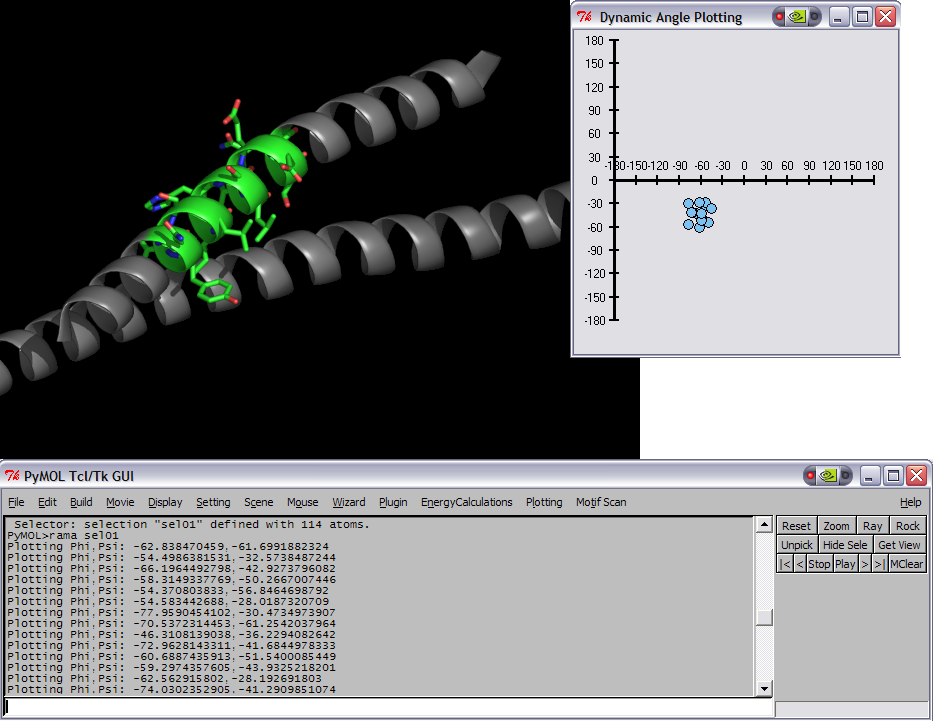
- RamaPlotBentComposite.png 935 × 720; 120 KB
- RamaPlotInitComposite.png 935 × 721; 133 KB
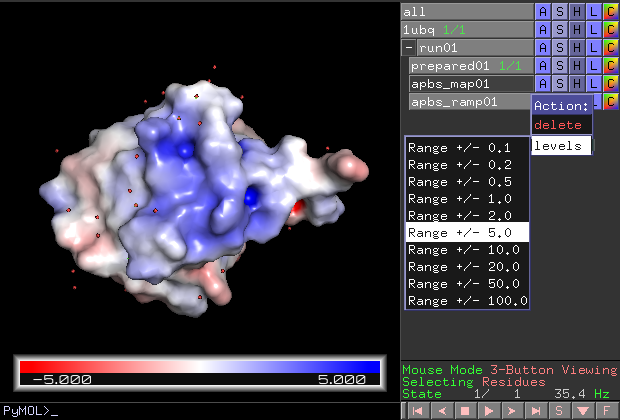
- Ramp-levels-menu.png 620 × 420; 88 KB
- Ray1.png 640 × 480; 218 KB
- Ray2.png 640 × 480; 210 KB
- Ray blend off.png 937 × 732; 240 KB
- Ray blend red.png 937 × 732; 227 KB
- Ray method off.png 955 × 756; 574 KB
- Ray method on.png 955 × 756; 575 KB