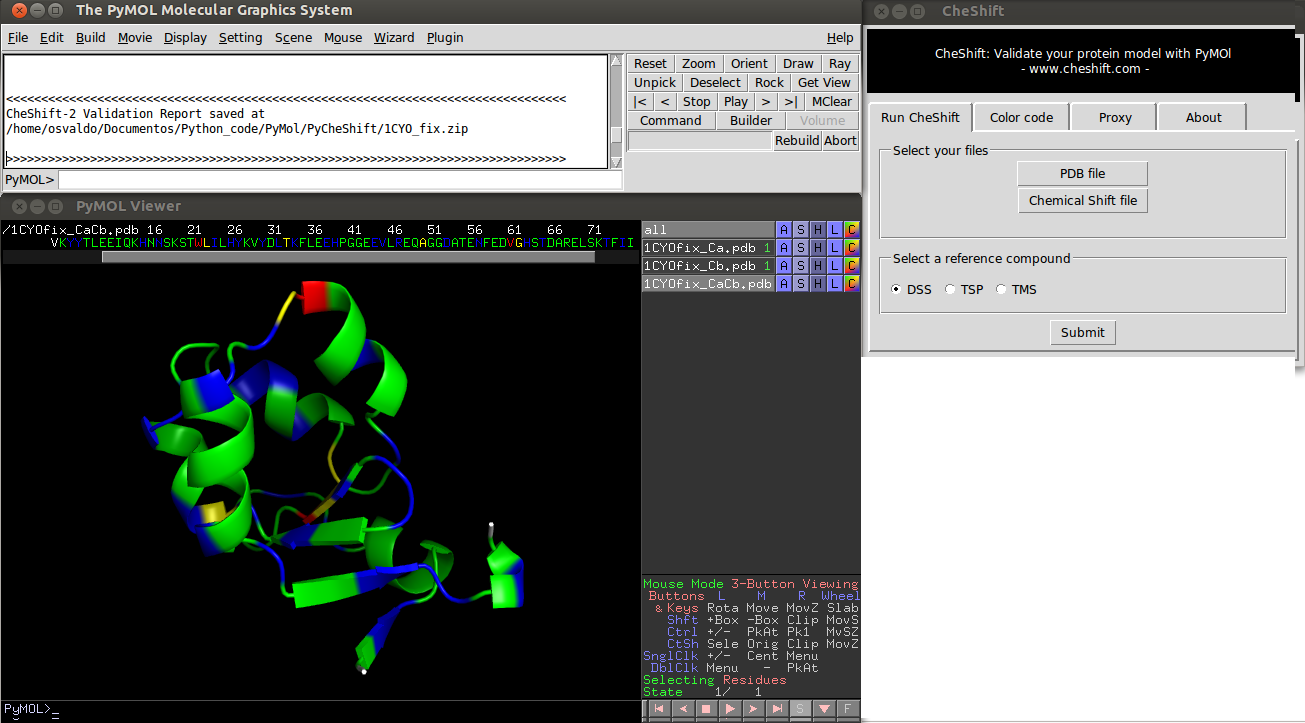
Result of a CheShift analysis.
Colors indicate the difference between predicted and observed
13C
α and/or
13C
βchemical shifts values averaged over all uploaded conformers. Green, yellow and red colors represent small, medium and large differences, respectively. White is used if either the prediction failed or the observed value is missing.
Blue is used to highlight residues for which the agreement between observed and predicted
13C
α and
13C
βchemical shifts can be improved, i.e., if the (χ
1 /χ
2 ) side-chain