This is a read-only mirror of pymolwiki.org
Talk:Label
Fixes
- Updates needed
New Page Content
New Page Overview
This is the content for the new labels page.
Old Page
DESCRIPTION
label allows one to configure the appearance of text labels for PyMOL objects. It labels one or more atoms properties over a selection using the python evaluator with a separate name space for each atom. The symbols defined in the name space are:
- name, the atom name
- resn, the residue name
- resi, the residue number/identifier
- chain, the chain name
- q,
- b, the occupancy/b-factor
- segi, the segment identifier
- type (ATOM,HETATM), the type of atom
- formal_charge, the formal charge
- partial_charge, the partial charge
- numeric_type, the numeric type
- text_type, the text type
All strings in the expression must be explicitly quoted. This operation typically takes several seconds per thousand atoms altered. To clear labels, simply omit the expression or set it to .
Label is great for labeling atoms, residues and objects. For a scene label, see Pseudoatom.
USAGE
label (selection),expression
SETTINGS
FONT
There are 10 different scalable fonts.
set label_font_id, number
where number is 5 through 14.
UTF8 Fonts
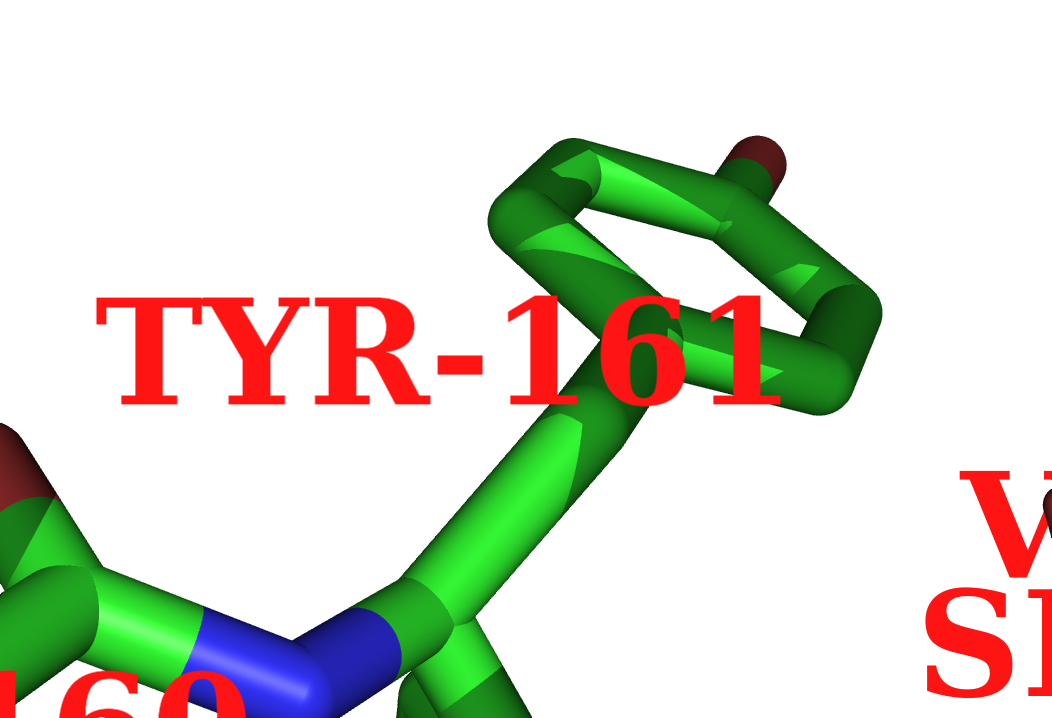
Newer versions support UTF8 fonts; use label_font_id from above to 15 or 16. The good news about the UTF8 fonts is that they support the alpha and beta characters. (See image.)
Here's some example code for the image at right:
# roman
set label_font_id, 15
set label_shadow_mode, 3
label 5/CA, "\316\261-Helix"
label 10/CA, "\316\262-Sheet"
# italic
set label_font_id, 16
# make bigger
set label_size, 50
Unicode Fonts
PyMOL gives you the flexibility to use encoded unicode fonts. This allows us to insert various symbols, like the symbol used for Angstrom. Here are the steps to insert a character from the unicode character set.
- Find the code for your character at Unicode Charts. The Angstrom character, <math>\textrm{\AA}</math> is u"\u00c5" and <math>\pm</math> is u"\u00b1".
- Label the selection. For simple strings, just type the string in double quote, -- "like this" -- and append to the end of that .encode('utf-8') -- "like this".encode('utf-8'). A working example is shown here,
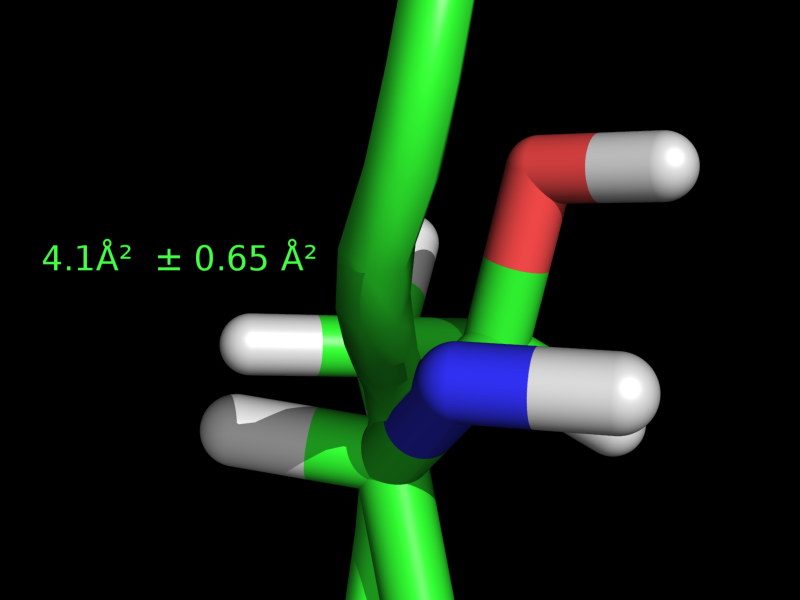
# label residue 30 with "4.1 Ang^2 +/- 0.65 Ang^2; see the image at right
label i. 30, "4.1" + u"\u00c5\u00b2 \u00b1 0.65 \u00c5\u00b2 ".encode('utf-8')
SIZE
The font size can be adjusted
set label_size, number
where number is the point size (or -number for Angstroms)
COLOR
Set a label's color by
set label_color, color
where color is a valid PyMol color.
If the coloring of the labels is not exactly the same as you'd expect (say black turns out grey, or red turns out pink), then try the following settings:
unset depth_cue
unset ray_label_specular
EXPRESSION
To set what the label reads (see above)
label selection, expression
For example
label all, name
label resi 10, b
POSITION
To position labels
edit_mode
then ctrl-middle-click-and-drag to position the label in space. (On Windows systems this appears to be shift-left-click-and-drag, presumably because those mice lack a true middle button.)
ctrl-shift-left-click-and-drag alters a label's z-plane. (Windows only? This may use the middle button, rather than shift-left, under *NIX / 3-button mice systems.)
EXAMPLES
Partial Charge
label (chain A),chain
label (n;ca),"%s-%s" % (resn,resi)
label (resi 200),"%1.3f" % partial_charge
Example 2
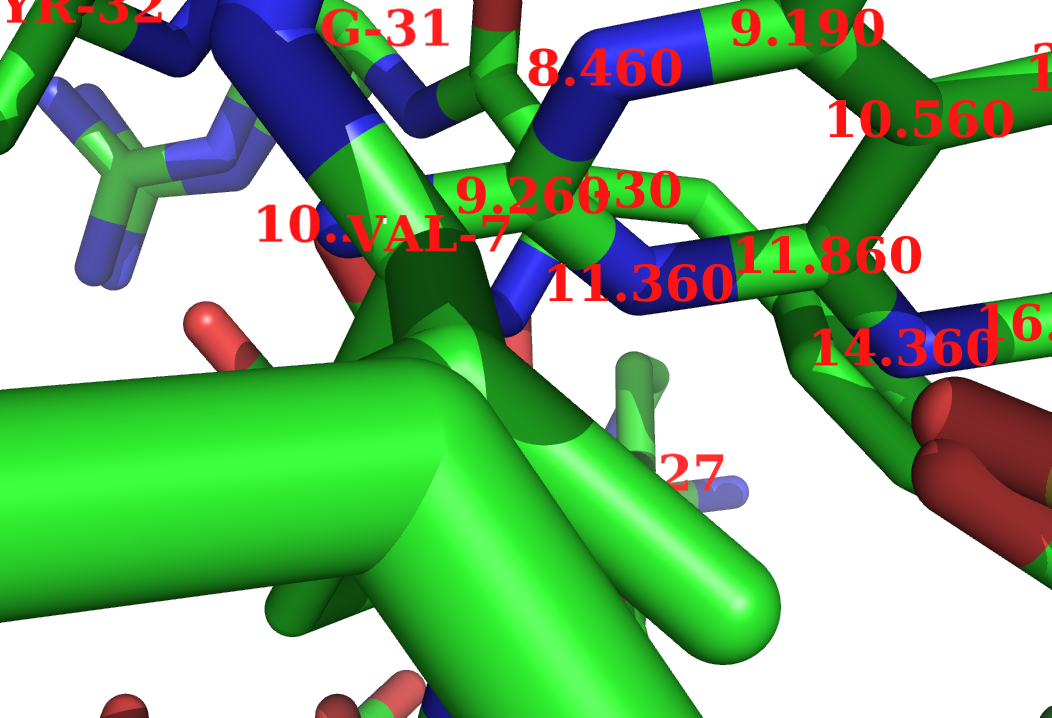
The following image was created with
label (resi 200),"%1.3f" % b
set label_font_id, 10
set label_size, 10
and finally, some labels were moved around in edit_mode.
More Advanced
This example shows how to label a selection with the XYZ coordinates of the atoms
from pymol import stored
stored.pos = []
# select the carbon atoms in my hetero atoms to label
select nn, het and e. C
# get the XYZ coordinates and put htem into stored.pos
iterate_state 1, (nn), stored.pos.append((x,y,z))
# label all N atoms. You need the pop() function or else
# PyMOL will complain b/c you didn't provide enough coords.
label nn, ("%5.5s, %5.5s, %5.5s") % stored.pos.pop()
Users Comments
Labels Using ID Numbers
The following commnent,
label SELECTION, " %s" % ID
labels the SELECTION with atom ID numbers.
You can make more complicated selections/lables such as
label SELECTION, " %s:%s %s" % (resi, resn, name)
which will give you something like "GLU:139 CG"
Labels Using One Letter Abbreviations
- First, Add this to your $HOME/.pymolrc file:
# start $HOME/.pymolrc modification
one_letter ={'VAL':'V', 'ILE':'I', 'LEU':'L', 'GLU':'E', 'GLN':'Q', \
'ASP':'D', 'ASN':'N', 'HIS':'H', 'TRP':'W', 'PHE':'F', 'TYR':'Y', \
'ARG':'R', 'LYS':'K', 'SER':'S', 'THR':'T', 'MET':'M', 'ALA':'A', \
'GLY':'G', 'PRO':'P', 'CYS':'C'}
# end modification
- . Second, instead of:
label n. ca, resn
use:
label n. ca, one_letter[resn]