This is a read-only mirror of pymolwiki.org
Difference between revisions of "Ccp4 contact"
m (Minor fixes) |
|||
| Line 1: | Line 1: | ||
| − | [[ | + | {{Infobox script-repo |
| + | |type = script | ||
| + | |filename = ccp4_contact.py | ||
| + | |author = [[Users:Dalyte|Gerhard]] | ||
| + | |license = GPL | ||
| + | }} | ||
== Overview == | == Overview == | ||
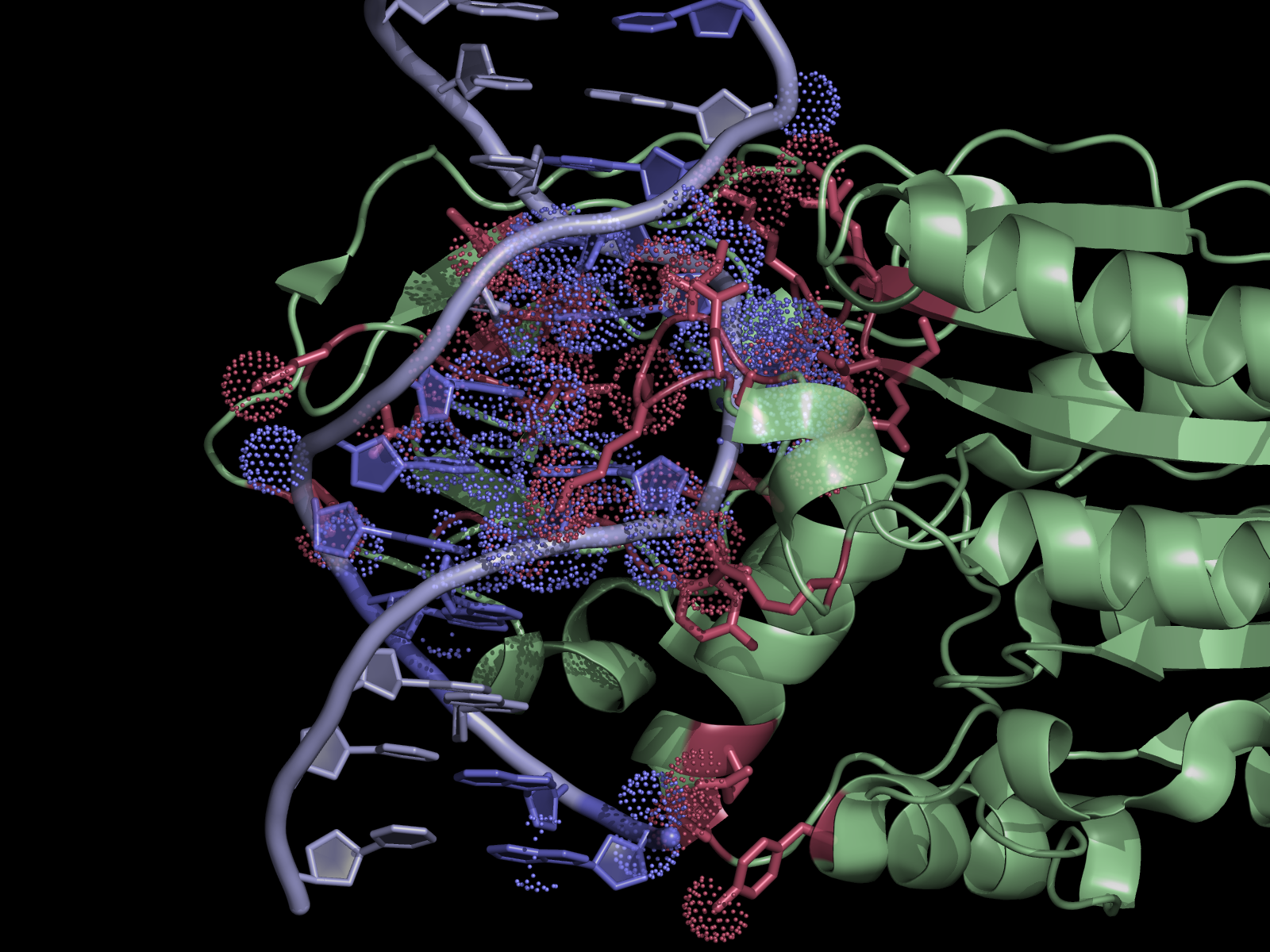
| + | [[File:HhaExample.png|thumb|300px|right|Interface residues (at cutoff <4A) in the 2c7r.pdb were found using NCONT, but similar results can be obtained using this script and CONTACT output. Usage of ccp4_contact script in PyMOL allows easy selection of residues and atoms listed in the output file. Interacting protein and DNA residues are colored in red and slate, respectively. Atoms in contact are shown in dots.]] | ||
The script selects residues and atoms from the list of the contacts found by CONTACT from CCP4 Program Suite (CONTACT analyses contacts between subsets of atoms in a PDB file). | The script selects residues and atoms from the list of the contacts found by CONTACT from CCP4 Program Suite (CONTACT analyses contacts between subsets of atoms in a PDB file). | ||
| − | First, we run CONTACT on our pdb file to find interface residues. Then by using the | + | First, we run CONTACT on our pdb file to find interface residues. Then by using the ccp4_contact script in PyMOL we separately select residues and atoms listed in the output file. This generates two selections (atoms and residues) for each interacting chain, allowing quick manipulation of (sometimes) extensive lists in CONTACT log file. |
== Usage == | == Usage == | ||
| − | + | ccp4_contact( contactsfile, selName1 = "source", selName2 = "target" ) | |
| − | |||
| − | |||
| − | |||
| + | == Example 1 == | ||
First use CONTACT to find interface residues/atoms in the pdb file. Once you have the log file proceed to PyMOL. | First use CONTACT to find interface residues/atoms in the pdb file. Once you have the log file proceed to PyMOL. | ||
| − | Make sure you | + | Make sure you import the ccp4_contact script first. |
fetch 2c7r | fetch 2c7r | ||
| − | selectCONTACTContacts | + | selectCONTACTContacts 2c7r.contact, selName1=prot, selName2=dna |
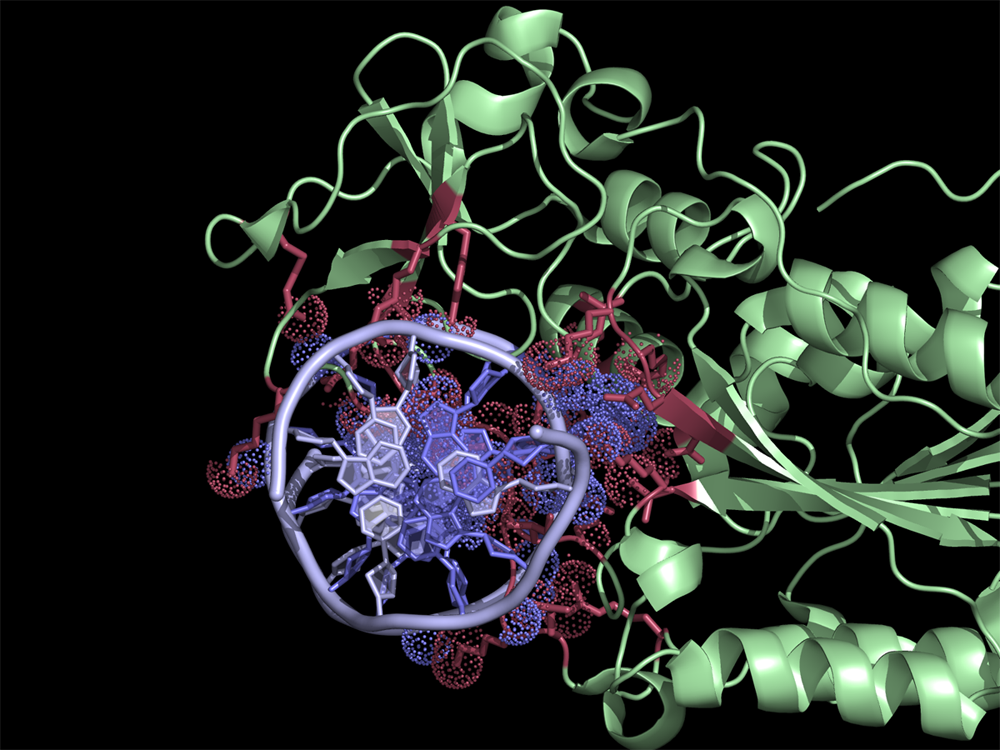
[[File:HhaI20example.png|thumb|300px|right|Quick and easy selection of interacting residues and atoms listed in the CONTACT log file. Protein and DNA residues are colored in red and slate, respectively. Atoms in contact are shown in dots.]] | [[File:HhaI20example.png|thumb|300px|right|Quick and easy selection of interacting residues and atoms listed in the CONTACT log file. Protein and DNA residues are colored in red and slate, respectively. Atoms in contact are shown in dots.]] | ||
| − | + | {{Template:PymolScriptRepoDownload|examples/ccp4_contact_1.pml}} | |
| − | + | <include src="https://raw.github.com/Pymol-Scripts/Pymol-script-repo/master/examples/ccp4_contact_1.pml" highlight="python" /> | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
[[Category:Script_Library]] [[Category:ThirdParty Scripts]] [[Category:Structural Biology Scripts]] | [[Category:Script_Library]] [[Category:ThirdParty Scripts]] [[Category:Structural Biology Scripts]] | ||
Revision as of 19:21, 14 December 2011
| Type | Python Script |
|---|---|
| Download | ccp4_contact.py |
| Author(s) | Gerhard |
| License | GPL |
| This code has been put under version control in the project Pymol-script-repo | |
Overview
The script selects residues and atoms from the list of the contacts found by CONTACT from CCP4 Program Suite (CONTACT analyses contacts between subsets of atoms in a PDB file). First, we run CONTACT on our pdb file to find interface residues. Then by using the ccp4_contact script in PyMOL we separately select residues and atoms listed in the output file. This generates two selections (atoms and residues) for each interacting chain, allowing quick manipulation of (sometimes) extensive lists in CONTACT log file.
Usage
ccp4_contact( contactsfile, selName1 = "source", selName2 = "target" )
Example 1
First use CONTACT to find interface residues/atoms in the pdb file. Once you have the log file proceed to PyMOL. Make sure you import the ccp4_contact script first.
fetch 2c7r selectCONTACTContacts 2c7r.contact, selName1=prot, selName2=dna
| Download: examples/ccp4_contact_1.pml | |
| This code has been put under version control in the project Pymol-script-repo | |
<include src="https://raw.github.com/Pymol-Scripts/Pymol-script-repo/master/examples/ccp4_contact_1.pml" highlight="python" />