This is a read-only mirror of pymolwiki.org
Dehydron
| Type | PyMOL Plugin |
|---|---|
| Download | plugins/dehydron.py |
| Author(s) | Osvaldo Martin |
| License | MIT |
| This code has been put under version control in the project Pymol-script-repo | |
Introduction
A dehydron is a protein backbone hydrogen bond incompletely shielded from water attack. A desolvated hydrogen bond is energetically more favourable than one exposed to the solvent and hence dehydrons are sticky , since they promote the removal of surrounding water through protein associations or ligand binding.
Dehydrons are less conserved than other structural motifs, hence identification of dehydrons could help to increase specificity during the rational drug design process. Certain proteins are enriched in dehydrons such as membrane proteins, toxic proteins and proteins that have a strong tendency to aggregate. Dehydrons have been invoked to explain biological processes above the molecular level such as the dosage imbalance effect in duplicated genes and the high connectivity of the protein interactomes of higher organisms.
A putative dehydron can be detected by counting the number of wrappers that surround a hydrogen bond. A wrapper is defined as a carbon atom not bonded directly to an oxygen or nitrogen atom, i.e. a non-polar carbon atom. A dehydron is defined by the number of wrappers inside two overlapping spheres centred at the Cα carbon of the donor and acceptor residues. If the number of wrappers around an hydrogen bond is below a certain cut-off value that hydrogen bond is identified as a dehydron.
Installation
Linux
This plugin is ready "out-of-box" for Linux users through the project Pymol-script-repo
Windows
This plugin is ready "out-of-box" for Windows users through the project Pymol-script-repo
Mac OsX
This plugin have not been tested on a Mac OsX machine, but it should work.
Usage
The plugin can be accessed using the following command:
wrappy [ selection [, angle_range [, max_distance [, desolv [, min_wrappers ]]]]]
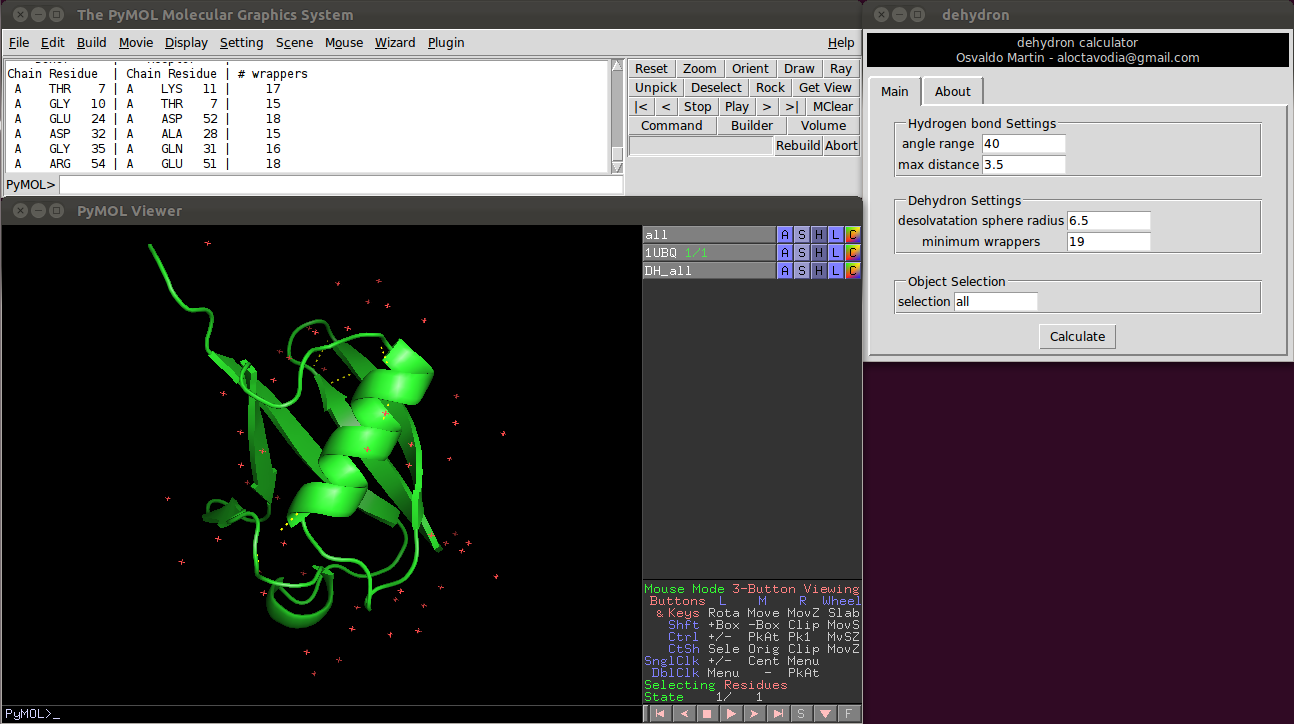
Or using a graphical environment (see figure 0)
There are five parameters the user can change:
Two of them control the hydrogen bonds detection.
- Angle range: deviation in degrees from the optimal hydrogen bond angle (C=0 N-H).
- Max distance: maximum donor-acceptor distance in angstroms.
Although no thoroughly optimized, the default values for the hydrogen bonds parameters were adjusted to get a close agreement between the hydrogen bond listed by wrappy and the ones listed by the What-If server.
Another two control the dehydron detection.
- Desolvatation sphere radius: this parameter controls the radius of the two spheres centred at the Cα carbon of the donor and acceptor residues. A dehydron is defined by the number of "wrappers" inside this two spheres.
- Min wrappers: a hydrogen bond surrounded with less "wrappers" than "min_wrappers" is a dehydron. Setting this parameter to a "high" value, something like 100, will return all main chain hydrogen bonds (according to the angle range and max distance parameters). The default value (19) was taken from a statistical analysis of ~7400 high-quality proteins form the Protein Data Bank (see below).
- Max wrappers: a hydrogen bond surrounded with more "wrappers" than "max_wrappers" is a buried hydrogen bond. The default value (35) was taken from a statistical analysis of ~7400 high-quality proteins form the Protein Data Bank (see below).
A wrapper is defined as a carbon atom not bonded directly to an oxygen or nitrogen atom, i.e. a non-polar carbon atom. The plug-in count as wrappers any non-polar carbon from any protein chain, organic ligand or other type of molecule, if the atoms belong "selection" (see below). This means that if you have, for example, a dimeric protein you will probably get different results for the dimer and for the isolated monomers. Instead, if you upload two (or more) different files the results will be independent because the plug-in does not count atoms from other objects
- Selection: This parameter allows the user to select which part of system is used to calculate dehydrons. This parameter is useful, for example, to calculate dehydronds for different objects independently or to easily calculate dehydrons with and without an organic ligand. The default selection is "all". Compute dehydrons for an specific selection is equivalent to delete all but the selected atoms and then compute dehydrons.
Methods
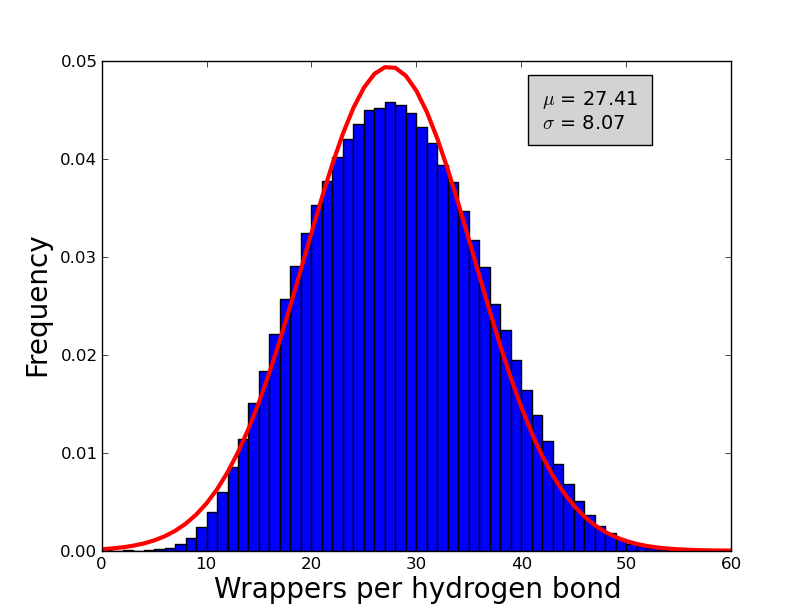
An analysis of 7476 high quality X-ray proteins was performed in order to estimate the number of wrappers that should be used as a cut-off to determine whether to call an hydrogen bond a dehydron (i.e. the min wrappers parameter). Although this value have been already estimated in the literature; differences in the algorithms used before and the ones used by wrappy could lead to differences in the exact value of the cut-off and hence this parameter was re-estimated in order to obtain reliable calculation of the dehydrons. To compute the numbers of wrappers the following parameters were used angle range = 40°, Max distance = 3.5 Å and desolvatation sphere = 6.5 Å. A non-redundant set of 7476 proteins were obtained from the Protein Data Bank. Each protein in this set conforms with the following criteria: Resolution < 2.0 Å, R-factors <= 0.25, not containing DNA and/or RNA molecules. Additionally, proteins with a sequence identity of 30% were removed. The frequency of wrappers in this set of proteins approximate a Gaussian distribution (see figure 1) with a mean of ~27 and a standard deviation of ~8, hence an hydrogen bond with less than 19 wrappers is defined as a dehydron (27-8 = 19). In the same fashion and over-wrapped hydrogen bond is and hydrogen bond with more than 35 wrappers (27+8).
Using the same set of 7476 proteins it was obtained that on average a high quality and globular protein should have 0.62 hydrogen bonds per residue (with a standard deviation of 0.06) and 17 wrappers per residue (with a standard deviation of 2). Hydrogen bonds per residue and wrappers per residue could be used as indicators of the global protein structure quality. Wrappy reports such indicators as z-score, i.e. the number of standard deviations an observation is above or below the expected mean value.
Acknowledgement
The H-bond detection code is based on the list_mc_hbonds.py script from Robert L. Campbell http://pldserver1.biochem.queensu.ca/~rlc/work/pymol/
Change log
- 2012-01-14 (Version 1.0)
- First public version was released and put under version control. In the project, Pymol-script-repo.
- 2012-01-28 (Version 1.1)
- Minor changes in the code most of them not visible for the end-user.
- 2012-02-28 (Version 1.5)
- The code was cleaned (e.g. remove global variables and other ugly stuff)
- The code was made available as a PyMOL command
- Better support for multiple objects
All features in this version and most of the code was provided by Thomas Holder, thanks Thomas! :-)
- 2012-03-14 (Version 1.6)
- Representation is not changed to "cartoon" after each calculation
- Total control over the selection from which dehydrons are calculated
- 2013-03-26 (Version 1.7)
- Wrappers were not correctly counted for structures with hydrogen atoms.
Thanks Shafqat Rasool for reporting the bug.
- 2013-07-19 (Version 2.0)
- The plug-in was renamed to wrappy.
- Wrappy reports, now, the hydrogen bonds per residue and the wrappers per residue as z-scores. The values of the mean and standard deviation, necessary to compute zscores, where taken from the analysis of ~7400 high quality X-ray proteins from the PDB.
- 2014-03-22 (Version 2.1)
- Wrappy reports all the hydrogens bonds. Dehydrons are displayed using red dashes, average wrapped hydrogens bonds are yellow, and over-wrapped hydrogens bonds are green.
References
Citation for Dehydrons:
De Simone, A., Dodson, G. G., Verma, C. S., Zagari, A., and Fraternali, F. (2005). Prion and water: tight and dynamical hydration sites have a key role in structural stability. Proceedings of the National Academy of Sciences of the United States of America, 102(21), 75357540.
Fernández, A. and Berry, R. S. (2003). Proteins with h-bond packing defects are highly interactive with lipid bilayers: Implications for amyloidogenesis. Proceedings of the National Academy of Sciences, 100(5), 2391–2396.
Fernández, A. and Crespo, A. (2008). Protein wrapping: a molecular marker for association, aggregation and drug design. Chemical Society Reviews, 37(11), 2373.
Fernández, A. and Lynch, M. (2011). Non-adaptive origins of interactome complexity. Nature, 474(7352), 502–505.
Fernández, A., Rogale, K., Scott, R., and Scheraga, H. A. (2004a). Inhibitor design by wrapping packing defects in HIV-1 proteins. Proceedings of the National Academy of Sciences of the United States of America, 101(32), 1164011645.
Fernández, A., Scott, R., and Berry, R. S. (2004b). The nonconserved wrapping of conserved protein folds reveals a trend toward increasing connectivity in proteomic networks. Proceedings of the National Academy of Sciences of the United States of America, 101(9), 28232827.
Kabsch, W. and Sander, C. (1983). Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers, 22(12), 2577–2637. PMID: 6667333.
Liang, H., Plazonic, K. R., Chen, J., Li, W.-H., and Fernández, A. (2008). Protein under-wrapping causes dosage sensitivity and decreases gene duplicability. PLoS Genetics, 4(1), e11.