This is a read-only mirror of pymolwiki.org
Dehydron
| Type | PyMOL Plugin |
|---|---|
| Download | plugins/dehydron.py |
| Author(s) | Osvaldo Martin |
| License | GPL |
| This code has been put under version control in the project Pymol-script-repo | |
Introduction
A dehydron calculator plugin for PyMOL. This plugin calculates dehydrons and display them onto the protein structure.
A dehydron is protein main chain hydrogen bond incompletely shielded from water attack. Dehydrons are sticky, since they promote the removal of surrounding water through protein associations or ligand binding. Dehydrons are less conserved than other structural motifs, hence identification of dehydrons could help to increase specificity during the rational drug design process.
For a brief introduction to the dehydron concept, please read wikipedia.
Installation
Linux
This plugin is ready "out-of-box" for Linux users through the project Pymol-script-repo
Windows
This plugin is ready "out-of-box" for win users through the project Pymol-script-repo
Usage
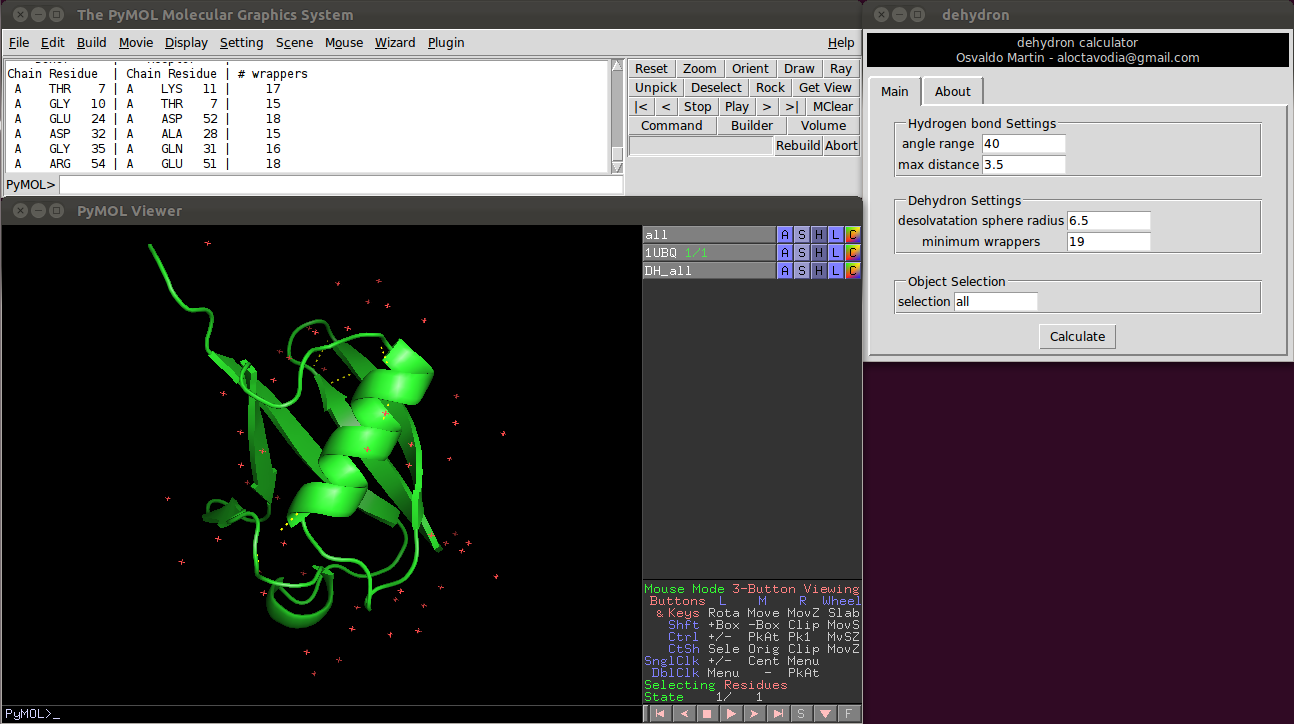
There are four parameters the user can change:
Two of them control the hydrogen bonds detection.
- Angle range: deviation in degrees from the optimal hydrogen bond angle (C=0 N-H).
- Max distance: maximum donor-aceptor distance.
And the other two control the dehydron detection
- Desolvatation sphere radius: this parameter controls the radius of the two spheres centered at the Cα carbon of the donor and acceptor residues. A dehydron is defined by the number of "wrappers" inside this two spheres.
- Cut-off wrappers: a hydrogen bond surrounded with less "wrappers" than "Cut-off wrappers" is a dehydron. Setting this parameter to a "high" value, something like 100, will return all main chain hydrogen bonds (according to the angle range and max distance parameters).
A wrapper is defined as a carbon atom not bonded directly to an oxygen or nitrogen atom, i.e. a non-polar carbon atom.
Acknowledgement
The H-bond detection code is based on the list_mc_hbonds.py script from Robert L. Campbell http://pldserver1.biochem.queensu.ca/~rlc/work/pymol/
References
Citation for Dehydrons:
Fernández A. and Scott R.; "Adherence of packing defects in soluble proteins", Phys. Rev. Lett. 91, 018102 (2003).
Fernández A., Rogale K., Scott R. and Scheraga H.A.; "Inhibitor design by wrapping packing defects in HIV-1 proteins", PNAS, 101, 11640-45 (2004).
Fernández A. "Transformative Concepts for Drug Design: Target Wrapping" (ISBN 978-3642117916), Springer-Verlag, Berlin, Heidelberg (2010).