This is a read-only mirror of pymolwiki.org
Dehydron
| Type | PyMOL Plugin |
|---|---|
| Download | plugins/dehydron.py |
| Author(s) | Osvaldo Martin |
| License | GPL |
| This code has been put under version control in the project Pymol-script-repo | |
Introduction
A dehydron calculator plugin for PyMOL. This plugin calculates dehydrons and display them onto the protein structure.
A dehydron is a main chain hydrogen bond incompletely shielded from water attack. For a brief introduction to the dehydron concept, please read http://en.wikipedia.org/wiki/Dehydron
Installation
Linux
This plugin is ready "out-of-box" for Linux users through the project Pymol-script-repo
Windows
This plugin is ready "out-of-box" for win users through the project Pymol-script-repo
Usage
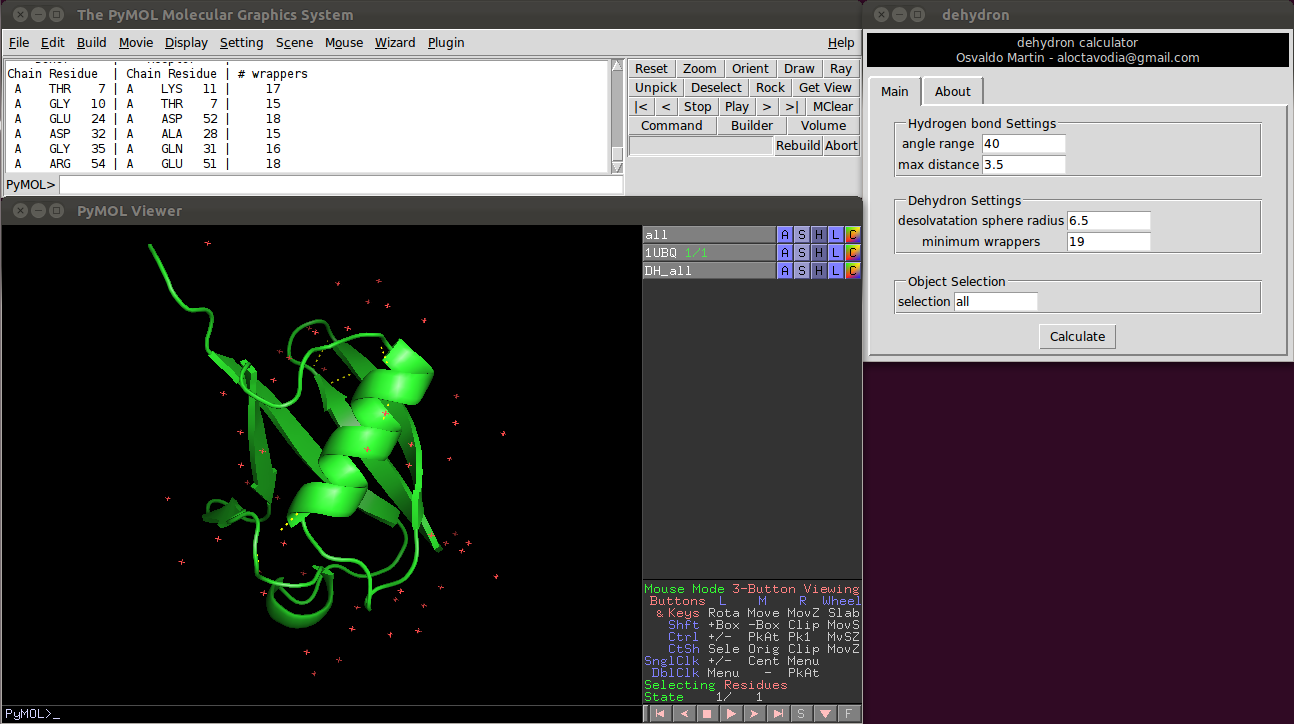
There are four parameters the user can change:
Two of them control the hydrogen bonds detection.
- Angle range: deviation in degrees from the optimal hydrogen bond angle (C=0 N-H).
- Max distance: maximum donor-aceptor distance.
And the other two control the dehydron detection
- Desolvatation sphere radius: this parameter controls the radius of the two spheres centered at the Cα carbon of the donor and acceptor residues. A dehydron is defined by the number of "wrappers" inside this two spheres.
- Cut-off wrappers: a hydrogen bond surrounded with less "wrappers" than "Cut-off wrappers" is a dehydron. Setting this parameter to a "high" value, something like 100, will return all main chain hydrogen bonds (according to the angle range and max distance parameters).
A wrapper is defined as a carbon atom not bonded directly to an oxygen or nitrogen atom, i.e. a non-polar carbon atom.
Acknowledgement
The H-bond detection code is based on the list_mc_hbonds.py script from Robert L. Campbell http://pldserver1.biochem.queensu.ca/~rlc/work/pymol/
References
Citation for Dehydrons:
Fernández A. and Scott R.; "Adherence of packing defects in soluble proteins", Phys. Rev. Lett. 91, 018102 (2003).
Fernández A., Rogale K., Scott R. and Scheraga H.A.; "Inhibitor design by wrapping packing defects in HIV-1 proteins", PNAS, 101, 11640-45 (2004).
Fernández A. "Transformative Concepts for Drug Design: Target Wrapping" (ISBN 978-3642117916), Springer-Verlag, Berlin, Heidelberg (2010).