This is a read-only mirror of pymolwiki.org
Difference between revisions of "DSSP Stride"
Jump to navigation
Jump to search
Hongbo zhu (talk | contribs) m |
Hongbo zhu (talk | contribs) m |
||
| Line 10: | Line 10: | ||
= News = | = News = | ||
| − | 2014_01_24_Update: The plugin is able to cope with empty chain name now. | + | * 2014_01_24_Update: The plugin is able to cope with empty chain name now. |
| − | 2012_10_22_Update: The most recent version of DSSP (v2.0.4) does not consider residues after the first [http://www.wwpdb.org/documentation/format33/sect9.html#TER <tt>TER</tt> record] from the same chain. The plugin has been updated to cope with this new feature. | + | |
| + | * 2012_10_22_Update: The most recent version of DSSP (v2.0.4) does not consider residues after the first [http://www.wwpdb.org/documentation/format33/sect9.html#TER <tt>TER</tt> record] from the same chain. The plugin has been updated to cope with this new feature. | ||
= External links = | = External links = | ||
Revision as of 08:34, 27 January 2014
Introduction
| Type | PyMOL Plugin |
|---|---|
| Download | plugins/dssp_stride.py |
| Author(s) | Hongbo Zhu |
| License | - |
| This code has been put under version control in the project Pymol-script-repo | |
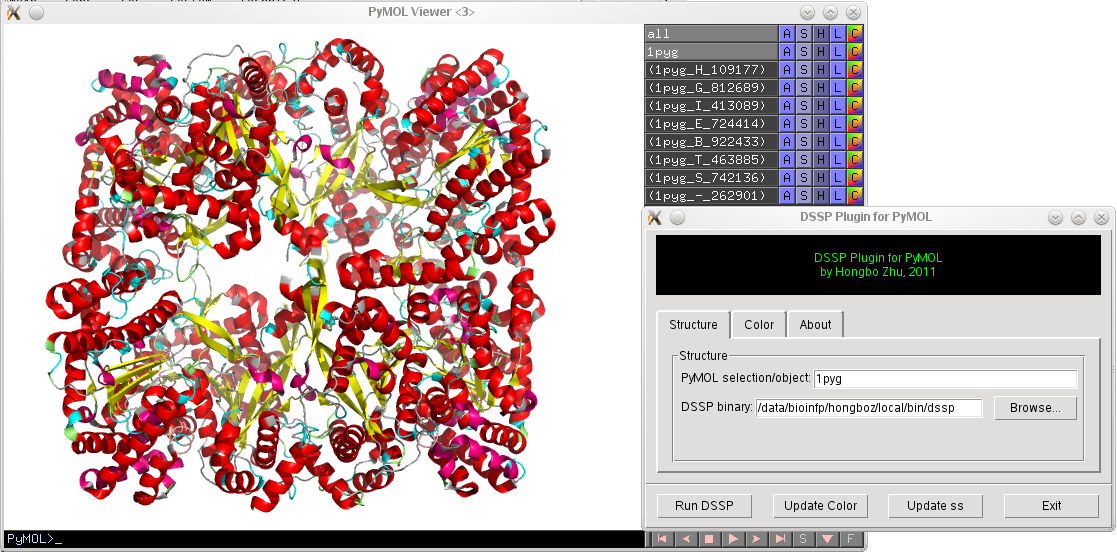
DSSP and Stride are popular tools for assigning secondary structures for proteins. DSSP & Stride Plugin for PyMOL provides a graphical user interface for coloring proteins according to their secondary structures assigned by DSSP or Stride. Most recent DSSP & Stride Plugin code can be obtained from this link.
News
- 2014_01_24_Update: The plugin is able to cope with empty chain name now.
- 2012_10_22_Update: The most recent version of DSSP (v2.0.4) does not consider residues after the first TER record from the same chain. The plugin has been updated to cope with this new feature.
External links
- Most recent code on github: DSSP & Stride Plugin for PyMOL @ GitHub
- Original site (not updated anymore): DSSP & Stride Plugin for PyMOL